Antimicrobial susceptibility tests: A comprehensive review of the most commonly used methods
Written by Éva Mészáros
13. August 2021
Microorganisms can stop responding to antimicrobial drugs when they acquire resistance, meaning that common infections and minor injuries that were once easy to treat can become life threatening. As of today, there are more than 700,000 deaths every year as a result of antimicrobial resistance, and this number could rise to 10 million by 2050 according to estimates by the WHO.1
Researchers and physicians worldwide are working every day to develop new antimicrobial drugs, detect existing resistances and find the best treatments for resistant infections in patients. In this article, we'll discuss the current situation with antimicrobials, explain how resistance develops, and take a closer look at the different antimicrobial susceptibility tests that are currently available.
Table of contents
1 The global threat of antimicrobial resistance
Antimicrobial resistance can develop naturally by random genetic mutations in pathogens, but the mis- and over-use of drugs – especially antibiotics in healthcare and agriculture – accelerate this process significantly. In countries where antibiotics are sold without a prescription, or used as growth-promoting substances or prophylactic additives in livestock farming, antibiotic resistant bacteria develop especially fast.2 Bacteria are able to pass on their resistance conferring genes quickly to other bacteria via horizontal gene transfer, and drug resistant pathogens can spread from animals to humans and between people, so it's only a matter of time before they reach every corner of the world.3
We could reduce the problem if we developed antimicrobial drugs at the same pace as resistant microorganisms emerge, but we're always a step behind. For example, the last new class of antibiotics was introduced in the 1980s and, since then, the clinical pipeline has been dry. The first challenge awaiting scientists searching for new antibiotics is the identification of antibacterial compounds. They usually look for them in nature, but some companies are now using artificial intelligence to trawl through chemical libraries to shortcut this process.4 Even after potential antibacterial compounds have been identified, many of them can't be pursued further, as less than 1 % are clinically useful due to efficacy problems – such as poor uptake by host cells – or toxicity issues.5
Until scientists develop new antimicrobials, physicians will have to rely on existing drugs, which will become more and more difficult as antimicrobial resistances evolve. Examples of multi- or even pan-resistant bacteria include Escherichia coli and Klebsiella pneumoniae strains that are common in the human intestinal tract. The rate of resistance to ciprofloxacin, which is used to treat urinary tract infections, is as high as 92.9 % for E. coli and 79.4 % for K. pneumoniae in some countries. Resistance to the alternative treatments – such as carbapenems for K. pneumoniae and fluoroquinolones for E. coli – is spreading as well. Therefore, colistin – an antibiotic that had gone out of use due to severe adverse effects, such as kidney damage – is often the last resort treatment. Unfortunately, some countries have already detected colistin-resistant bacteria for which there is no other effective antibiotic treatment at present.6,7
2 How to perform antimicrobial susceptibility tests
There are several methods available for when physicians need to know which drug is suitable to treat a patient's infection, or when scientists want to determine the efficacy of a new antimicrobial compound. In the following sections, we will take a closer look at the most common ones.
2.1 Determination of minimum inhibitory concentration
One way to measure the antimicrobial susceptibility of a pathogen is to determine the minimum inhibitory concentration (MIC). The MIC is defined as the lowest concentration of an antimicrobial drug that inhibits the growth of a microorganism. It can either be determined using broth dilution, agar dilution or antimicrobial gradient methods.
2.1.1 Broth dilution method
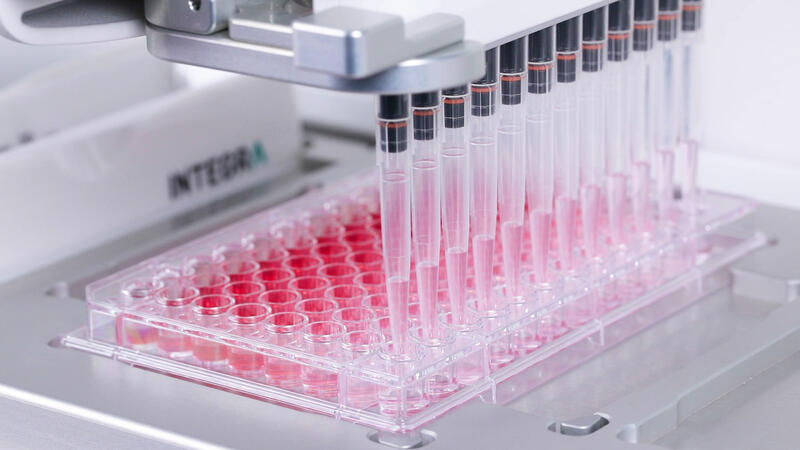
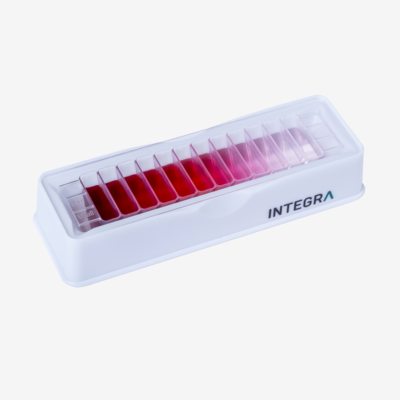
For the broth dilution technique, you create a two-fold serial dilution of an antimicrobial compound in liquid growth medium, and inoculate it with a predefined amount of a pathogenic microorganism. Previously, scientists used to perform this assay in tubes (macrobroth method) but, today, 96 well plates are commonly used (microbroth method). After incubation of the inoculated 96 well plate, a microplate reader measures the microbial growth to determine the lowest effective concentration of the antimicrobial compound. To see an example of a microbroth protocol, see our application note on Automated Minimum Inhibitory Concentration Testing.
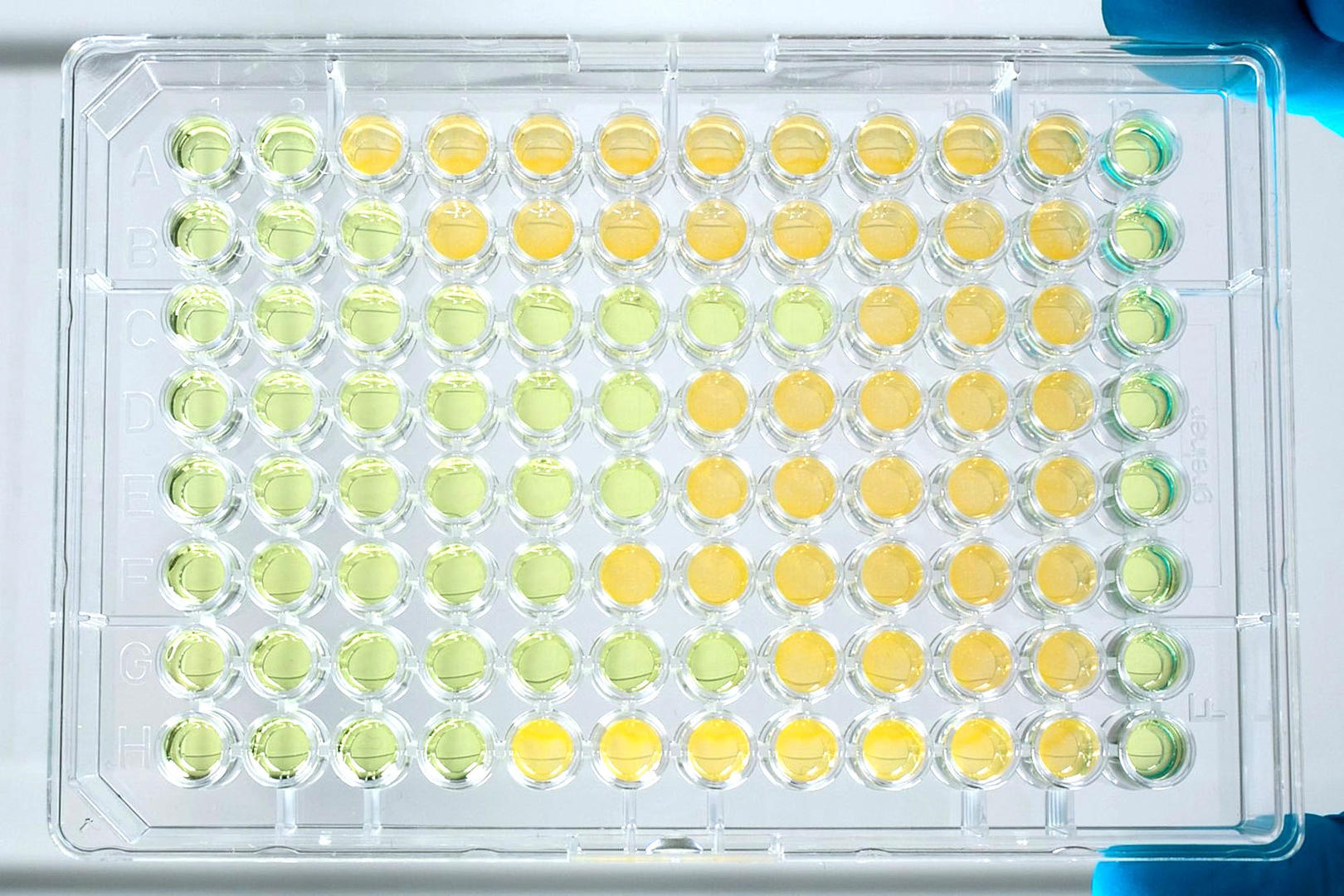

If you want to identify not only the MIC, but also the lowest concentration resulting in microbial death (MMC = minimum microbicidal concentration), you can subculture the broth dilution of your MIC tests to agar plates that don't contain the antimicrobial compound. If the pathogenic microorganisms start to proliferate, the antimicrobial didn't cause their death but only inhibited their growth.
An advantage of the broth dilution method is that it provides a quantitative result. However, as antimicrobials are usually tested in two-fold serial dilutions, you can’t measure the 'true' MIC, which lies somewhere in between the lowest test concentration that inhibits microbial growth and the highest test concentration that doesn't inhibit microbial growth.8 Another plus is that it's a space saving method, as you can test up to 12 compounds in a single microplate.9 The major downside is that the preparation of serial dilutions involves a large number of repetitive pipetting steps which are tedious, time consuming and error and contamination prone if performed manually. The broth dilution method is therefore mostly used in research, and high throughput labs often opt for a walk-away pipetting solution for serial dilutions.
2.1.1.1 Automated broth-based susceptibility testing
Labs dedicating a large amount of their time to MIC testing might even consider investing in a fully automated susceptibility testing system, such as MicroScan WalkAway, Phoenix™, Vitek 2 or Sensititre™ ARIS™ 2X. For most of these systems, you need 96 well plates prefilled with dehydrated reagents and antimicrobials, which you hydrate and inoculate manually or with an autoinoculator. With the Vitek 2, this step can be skipped, as it uses compact plastic reagent cards in a 64 well format that it can fill itself. The inoculated 96 well plates or reagent cards are then incubated by the systems, and analyzed photometrically, fluorometrically and/or colorimetrically.9,10
As automated systems periodically measure microbial growth during incubation, they can usually provide susceptibility test results in a shorter period of time compared to manual broth dilution assays.9,10 However, the instruments and consumables are more expensive and, as you need to purchase 96 well plates or reagent cards containing the antimicrobials you want to test, they are only suited for tests with known antimicrobials, and not for drug discovery assays where new compounds are assessed.
2.1.2 Agar dilution method
Another MIC determination method is agar dilution. Instead of creating the two-fold serial dilution of the antimicrobial compound in liquid growth medium, you add different concentrations of it into agar medium samples that are then poured into Petri dishes. Once the agar is set and dried, you can plate out up to 60 microbial inocula in one dish. After incubation of the dishes, you determine the MIC by identifying the lowest antimicrobial concentration that completely inhibits visible microbial growth.8,11,12
Agar dilution is a great method to test multiple microbial pathogens against a single compound, and is also recommended for fastidious organisms, such as anaerobes and Helicobacter species.11 If not semi-automated with an inoculum replicator, it can be a very time-consuming method and, similar to the broth dilution assays, the real MIC value lies somewhere between the lowest concentration inhibiting growth and the next lower one tested.
2.1.3 Antimicrobial gradient method (Etest®)
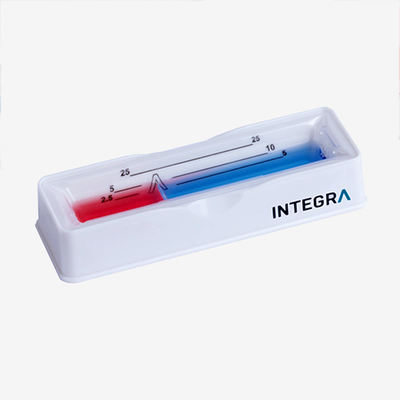
The antimicrobial gradient method is also performed in Petri dishes filled with agar, but in contrast to the agar dilution method, it's used to test various compounds simultaneously. The first step is the inoculation of an agar plate with the microorganisms of interest. Etests from bioMérieux – which have under surfaces impregnated with a concentration gradient of a certain antimicrobial compound and upper surfaces that show a concentration scale – are then placed on the agar. During incubation, the antimicrobial diffuses into the media, and a growth inhibition ellipse forms. The MIC of the antimicrobial compound corresponds to the value on the concentration scale that is located where the ellipse and the strip intersect.9,11
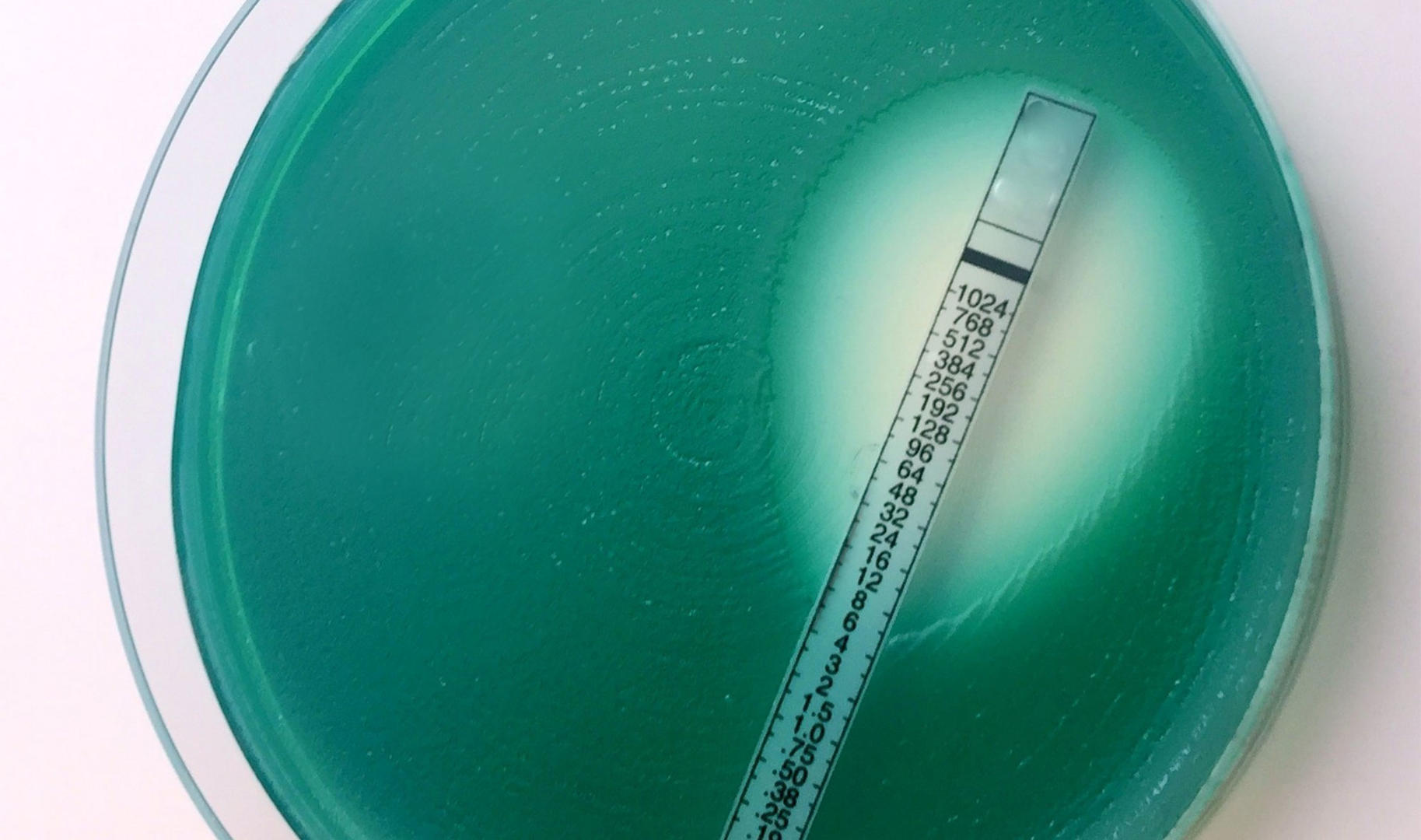

Etests are a very simple method to determine the MIC, and generally provide results correlating well with both broth and agar dilution assays. However, if a large number of antimicrobial compounds has to be tested, it can be quite costly to buy the strips, which is why Etests are usually only used for low throughput assays, or when a fastidious organism requiring enriched culture medium or a special incubation atmosphere is being tested.9,11 On top of that, Etests can only be used for tests with known antimicrobials, as strips impregnated with candidate compounds aren't available.
2.2 Disk diffusion method
The disk diffusion method is similar to the antimicrobial gradient method but provides a qualitative result. Instead of placing Etest strips on the inoculated agar, up to 12 small paper disks containing specified concentrations of different antimicrobial compounds are used. The antimicrobial agents will set up concentration gradients when diffusing into the culture media and, during the subsequent incubation step, microbes will grow across the plate except in the so-called 'zones of inhibition' around the paper disks where the antibiotic concentration is high enough to inhibit their growth. If all the paper disks in the plate have the same antimicrobial concentration and diffusibility, it can be concluded that the one with the largest zone of inhibition is the most effective.8,9
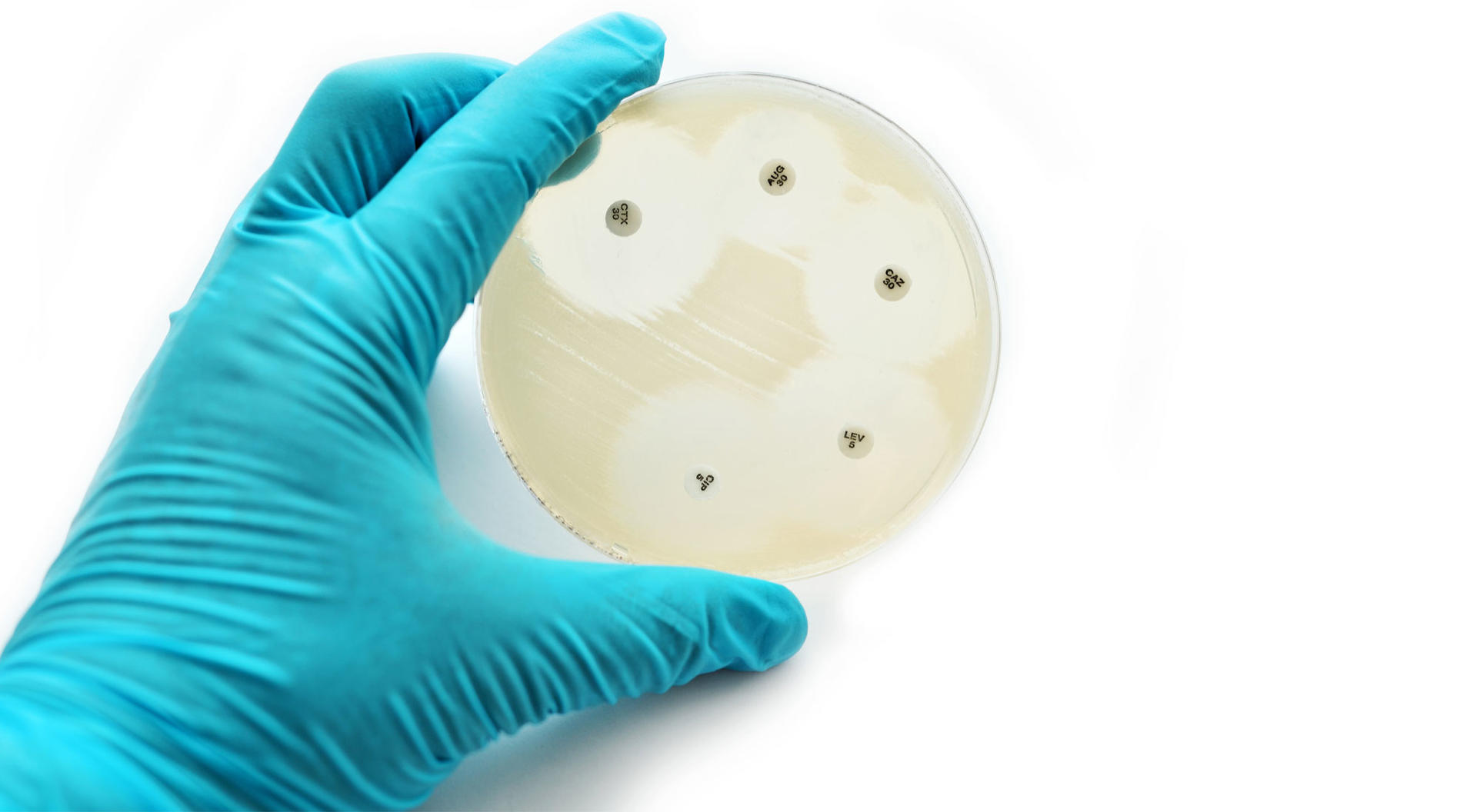
The disk diffusion method is very affordable, but needs to be performed manually. On top of that, it only provides a qualitative result, unless you're using a zone reader system capable of calculating an approximate MIC for some microorganism-antimicrobial combinations using an algorithm.8,9 Another downside is that you can't use it to test new antimicrobial compounds, as disks are only available for currently used antimicrobial drugs.
2.3 Genetic methods for assessing antimicrobial resistance
In addition to the phenotypic methods described above, there are various genotypic methods. These generally start with PCR to amplify the DNA sequence of interest from the pathogen. Various molecular or genomic DNA-, amplification- or sequencing-based methods are available to determine if the pathogen has specific resistance genes or genetic mutations, including DNA microarrays, electrophoresis, Southern blotting, restriction fragment-length polymorphism, single-strand conformation polymorphism, DNA fingerprinting, molecular beacons and DNA sequencing.13,14
When working with patient samples, genotypic methods are generally much faster than phenotypic methods, as you don't need to isolate and culture the microbial pathogen of interest before starting your susceptibility tests. The absence of a microbial culture not only saves time – especially when working with difficult-to-cultivate or slow-growing microbes – but also reduces the biohazard risk to laboratory personnel. The major disadvantage of genotypic methods is that you need to know the exact genetic sequence or gene to test for. You can only use these techniques if the genetic mechanism of resistance has been defined, and if a suitable detection assay has been developed. As a result, new forms of resistance can only be detected using culture-based phenotypic methods.13,15
3 Conclusion
Whether the WHO estimates of 10 million deaths due to antimicrobial resistance in 2050 will come true depends, among many other factors, on whether new antimicrobial drugs come onto the market in the next decades. We hope that we have been able to make a small contribution to the discovery of new compounds with this article, by helping you to choose the right antimicrobial susceptibility method.
Did you like this blog post? Then sign up for our newsletter: