Simple PCR tips that can make or break your success
Written by Marlene Frank
23. June 2022
Since the outbreak of the COVID-19 pandemic, PCR is on everybody's lips. However, only people working in the lab know how difficult it can be to get the desired results using this well-established technique. Out of this frustration came the popular joke that PCR should stand for ’pipette, cry, repeat’. To ensure that this stays a joke from now on, and that your PCR reactions never drive you to despair again, we have compiled the most important tips and tricks for a successful PCR set-up.
Table of contents
What is PCR?
The polymerase chain reaction (PCR) is used to amplify specific DNA sequences for downstream use. Its inventor Kary Mullis, whose patent on PCR was approved in 1987, was awarded the Nobel Prize in Chemistry six years later,1 and since this time, PCR has remained one of the most essential molecular biology techniques. Genetic engineering, genotyping, sequencing and the identification of familial relationships, to name a few examples, wouldn't be possible without it.
To perform PCR reactions, you need to prepare a master mix, add template DNA, and amplify the sequence of interest using a thermal cycler. If you want to learn what the components of the master mix are, and how they interact with the template DNA during the thermal cycles, read our blog post The complete guide to PCR.
PCR tips and tricks
Setting up a PCR reaction might seem straightforward, but it is far from it. Calculating the required amounts of master mix reagents correctly to get the right volume, at the right concentration, is the first challenge.
Once this is accomplished, the reagents need to be mixed together. The difficulty here is that the liquids usually have to be cooled and they are often highly viscous, sticky and needed in minimal quantities. In addition, work must be performed in a concentrated manner, as distractions or interruptions can quickly lead to a situation where you no longer know which reagents have already been added to the master mix. Errors such as skipping a tube or well can also easily occur when filling PCR strips or plates with master mix and adding template DNA, especially when using single channel pipettes.
The last and probably biggest challenge is to keep your PCR reactions free from contamination. PCR is a very sensitive assay that can create a large number of nucleic acid copies from a tiny amount of starting material, so amplicon and sample contamination can be a huge problem.
Master mix calculations
Let's first have a look at the mathematical calculations needed to set up a PCR master mix. We'll assume that you want to set up several PCR reactions with a volume of 50 µl each.
To calculate the required volume for each reagent, it is best to create a table (see Table 1) with the necessary components, and fill in the stock concentrations and desired final concentrations for the buffer, the MgCl2, the dNTPs and the primers. Then, calculate the dilution factors by dividing the stock concentration by the final concentration. To determine the volume needed for a single PCR reaction, divide the desired reaction volume by the dilution factor.2
For the polymerase, a slightly different equation is needed. The manufacturer of the enzyme will tell you the amount of polymerase in one µl, e.g. 5 Units/µl. Fill in this value in the column for the stock concentration and put the desired amount – e.g. 1.25 Units – in the column for the final concentration. The volume needed can then be calculated as follows: 1.25 Units x (1 µl / 5 Units) = 0.25 µl.3
The template DNA volume required depends on your sample type. You should add about 1 pg to 10 ng of plasmid or viral DNA, and 1 ng to 1 µg of genomic DNA. In the example below, we calculated how much you would need to use for 0.5 µl of a 1 µg/µl template DNA.4
Finally, add the required volumes for all the reagents. The difference between the desired total reaction volume (50 µl) and the result obtained gives you the volume of PCR-grade water.5
| Reagent | Stock conc. | Final conc. (CF) |
Dilution factor (= stock con. / CF) |
Volume needed (= 50 µl / dil. factor) |
|---|---|---|---|---|
| Buffer | 10X | 1X | 10 | 5 µl |
| MgCl2 | 25 mM | 1.5 mM | 16.66 | 3 µl |
| dNTPs | 10 mM | 0.2 mM | 50 | 1 µl |
| Forward primer | 10 µM | 250 nM | 40 | 1.25 µl |
| Reverse primer | 10 µM | 250 nM | 40 | 1.25 µl |
| Polymerase | 5 Units/µl | 1.25 Units | - | 0.25 µl |
| Template DNA | 1 µg/µl | - | - | 0.5 µl |
| PCR-grade water | - | - | - | 37.75 µl |
Table 1: Example of a PCR master mix table
After determining the required reagent volumes for one PCR reaction, you can simply multiply them by your sample number (plus the negative and positive controls) to get the total volumes for the entire PCR set-up. We recommend adding one additional aliquot to that result, as some of the master mix may be lost during pipetting due to evaporation, adherence to the tip, or pipetting inaccuracies.
That's it, you are now ready to set up your PCR reactions by following the best pipetting practices listed below.
Best PCR pipetting practices
Start by preparing your master mix from all the components listed above, except the template DNA. The huge advantage of preparing the entire quantity of master mix needed for an experiment, and subsequently transferring single aliquots into PCR strips or plates, is that you can pipette higher volumes with better accuracy. On top of that, it reduces pipetting steps, making the entire process less tiring and error prone. Since pipetting mistakes cannot be completely ruled out, you should add the master mix components in order of their price, starting with the most affordable reagent. This way, you waste less money if you have to start over.6
Once your master mix is finished, well mixed and dispensed into tubes or plates, you can add the template DNA. As the DNA samples are usually highly viscous and needed in small quantities, you should either dispense them into the master mix or onto the wall of the tube or well. After dispensing, keep the plunger depressed while dragging the tip gently along the wall of your labware to remove any residual liquid. In addition, we recommend using low retention tips.
If you're not using a hot-start polymerase, cool your reagents throughout the entire process of master mix preparation and sample addition, to prevent non-specific amplification.
When you are ready to load your samples into the thermal cycler, check that they are tightly capped or sealed, and spin them down to ensure that no droplets remain on the labware wall during amplification.
Pipetting solutions for PCR reactions
Before discussing various pipetting solutions, we would like to address one of the most important aspects of liquid handling. No matter which pipettes you choose, ensure that they are well maintained by regularly calibrating them and checking their performance in between uses.
The most affordable pipettes for master mix preparation would be manual single channel models. However, as you need to accurately measure and mix several very expensive reagents, we recommend investing in electronic single channel pipettes. The motor-controlled piston movement guarantees that they always dispense the exact desired volume, minimizing variability to increase the precision and accuracy of pipetting.
For the container, you can either prepare the master mix in a tube or, if you intend to transfer it with an electronic multichannel pipette, in a low dead volume reagent reservoir. The combination of an electronic multichannel pipette and a reservoir is ideal for this step, because you can fill several tubes or wells simultaneously. On top of that, electronic multichannel pipettes usually feature a repeat dispense mode, allowing you to aspirate a large volume of master mix, then dispense it into multiple smaller aliquots. It is also possible to use an electronic single channel pipette if you have a low throughput.
To add template DNA to the master mix aliquots, an adjustable tip spacing pipette can be very handy if the labware format of your samples doesn't match the container used for PCR amplification. For example, it allows you to transfer several template DNA samples from microcentrifuge tubes to an entire row or column of a 96 well plate in one step.
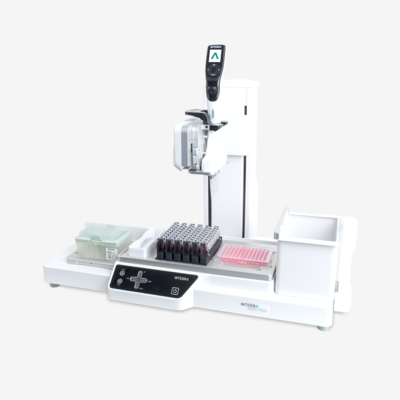
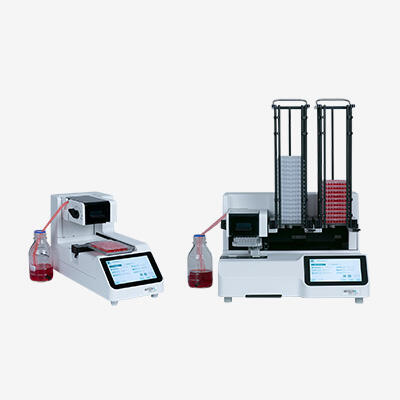
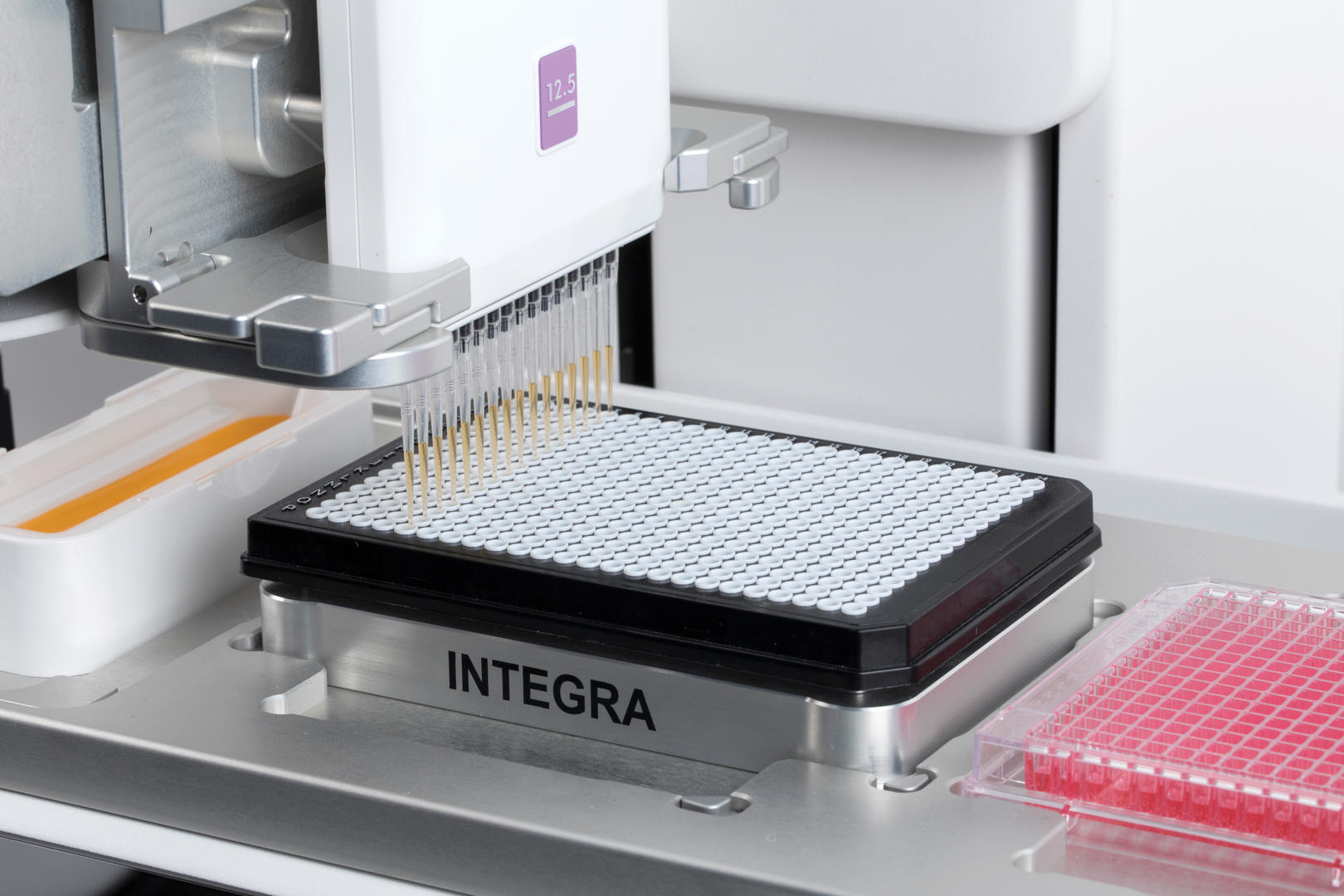
High throughput labs might even want to take advantage of automated solutions for master mix plating and sample transfer, such as a pipetting platform that is capable of automating electronic pipettes.

How to prevent PCR contamination
Several preventative measures should be taken to avoid contaminating your master mix or template DNA with amplicons that were generated during previous PCRs.
One of the most effective means is to physically separate the master mix preparation, template DNA addition, amplification and analysis areas from one another, and to work in laminar flow or biosafety cabinets. Each work zone, and its corresponding equipment, should be cleaned before and after an experiment, and tools used in one area should never enter another one. Learn more on the ideal set-up of a PCR lab.
When it comes to consumables, make sure you purchase sterile products that are certified to be free from DNase, RNase and PCR inhibitors. Pipette tips should form a perfect seal with the pipette to eliminate contamination that may occur when tips drip or fall off. Using filter tips will also avoid the risk of aerosols entering your pipettes and contaminating subsequent PCR reactions.
As you're a potential source of contamination too, always wear gloves to prevent introducing enzymes, microbes and skin cells to the reaction, and change them when going from one area to another. On top of that, keep your tubes closed whenever possible during the entire PCR set-up.
Despite these preventative measures, you can't completely eliminate the possibility of contaminated PCR reactions. To avoid having to throw away your entire stock of a certain reagent if this occurs, prepare single use aliquots of your master mix components. You can also prepare aliquots of positive and negative controls, as well as serial dilutions of standards for quantitative PCR (qPCR) assays, ahead of time. Electronic pipettes with repeat dispense and serial dilute modes can be helpful for this task, not only to reduce the risk of contamination, but also to increase the efficiency of PCR set-up.
Conclusion
PCR is a fundamental technique in research, diagnostics and forensics. It often involves pipetting minuscule reagent volumes with tricky properties, so it can be difficult to obtain the desired results. On top of that, contamination can have a huge impact on results, as it's a very sensitive assay. We hope that the tips and tricks provided in this article will help you make your future PCR reactions a success. Many of these recommendations can also be applied to other amplification assays, such as reverse transcription and qPCR, loop-mediated isothermal amplification (LAMP) and helicase-dependent amplification (HDA).
Did you like this article?
Further reading: