Enzyme essentials: a comprehensive guide to the catalysts of life
Written by Anina Werner
03. April 2024
This comprehensive blog shedding light on the world of enzymes explores how these tiny catalysts play an indispensable role in essential biochemical reactions. We will describe their structure, explain how they work, and uncover the intricacies of enzyme catalysis. Furthermore, we will explore the factors influencing enzyme activity, such as temperature, pH, and substrate concentration, and discuss the regulation of enzyme action through inhibition and activation. Read on to learn everything you need to know about enzymes, the different types and their functions.
Table of contents
What are enzymes?
Most enzymes are proteins made of chemically linked amino acid monomers, although some are ribonucleic acid (RNA) molecules. Enzymes can be found in all living organisms and play a key role in speeding up or catalyzing biochemical reactions – such as metabolic processes – by lowering the activation energy needed to start the reactions. Without enzymes, most vital chemical processes would occur too slowly to sustain life. An extreme example of a catalyst is the enzyme orotidine 5'-phosphate decarboxylase, which catalyzes a reaction in 18 milliseconds that, without its presence, is estimated to take 78 million years.1
How do enzymes work?
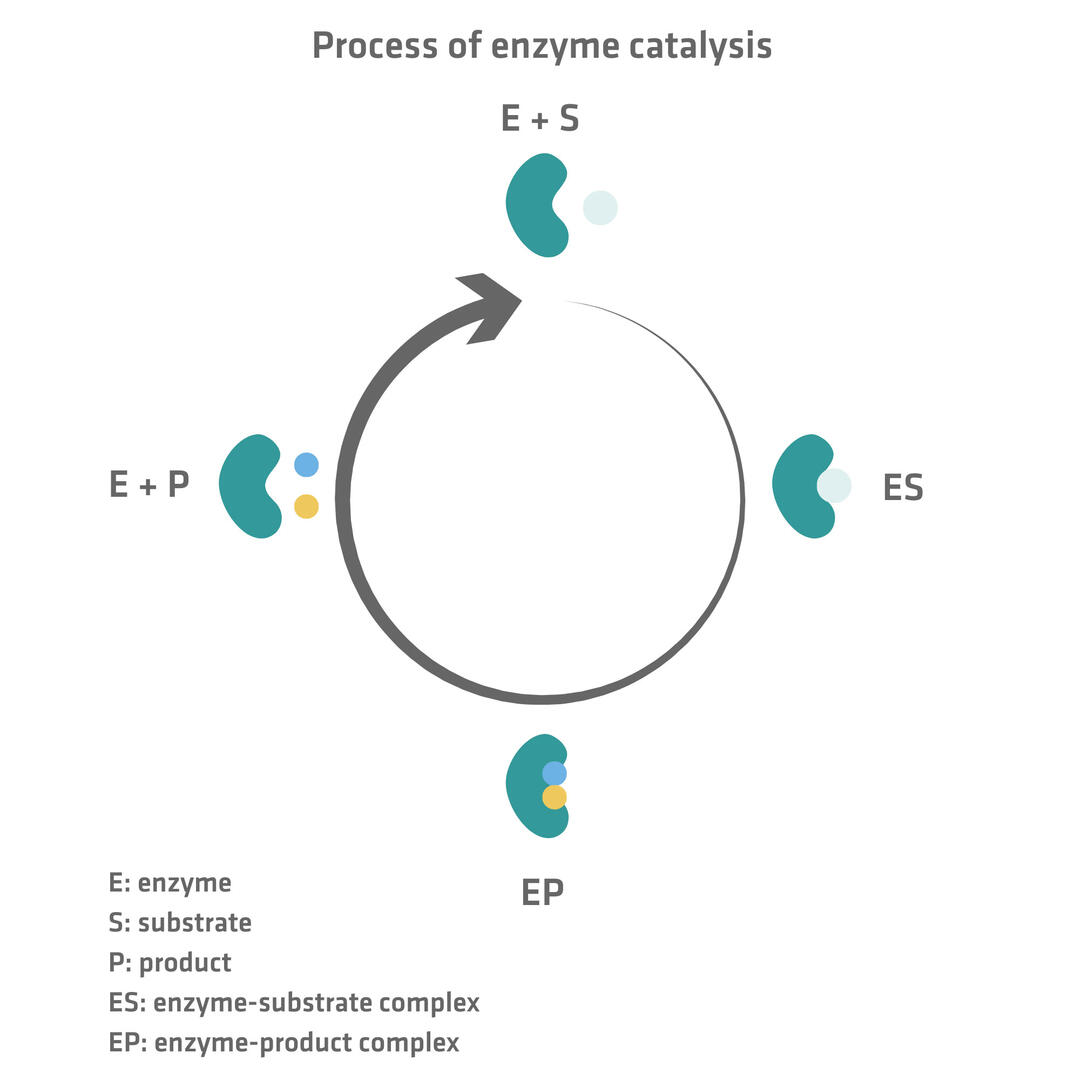
Enzymes bind to molecules called substrates to catalyze biochemical reactions in cells. Once the two form a complex, the enzyme converts the substrate into another distinct molecule called a product and releases it. Enzymes are not consumed in this process, so can perform the same reaction repeatedly.
Structure of enzymes
To better understand the enzyme catalysis process outlined above, let's have a look at the structure of enzymes. Enzymes have several different parts:
- One or more polypeptide chains made of chemically linked amino acids: these linear chains usually consist of 62 to 2500 amino acid residues,2 and the amino acid sequence determines the 3D structure of the enzyme.3
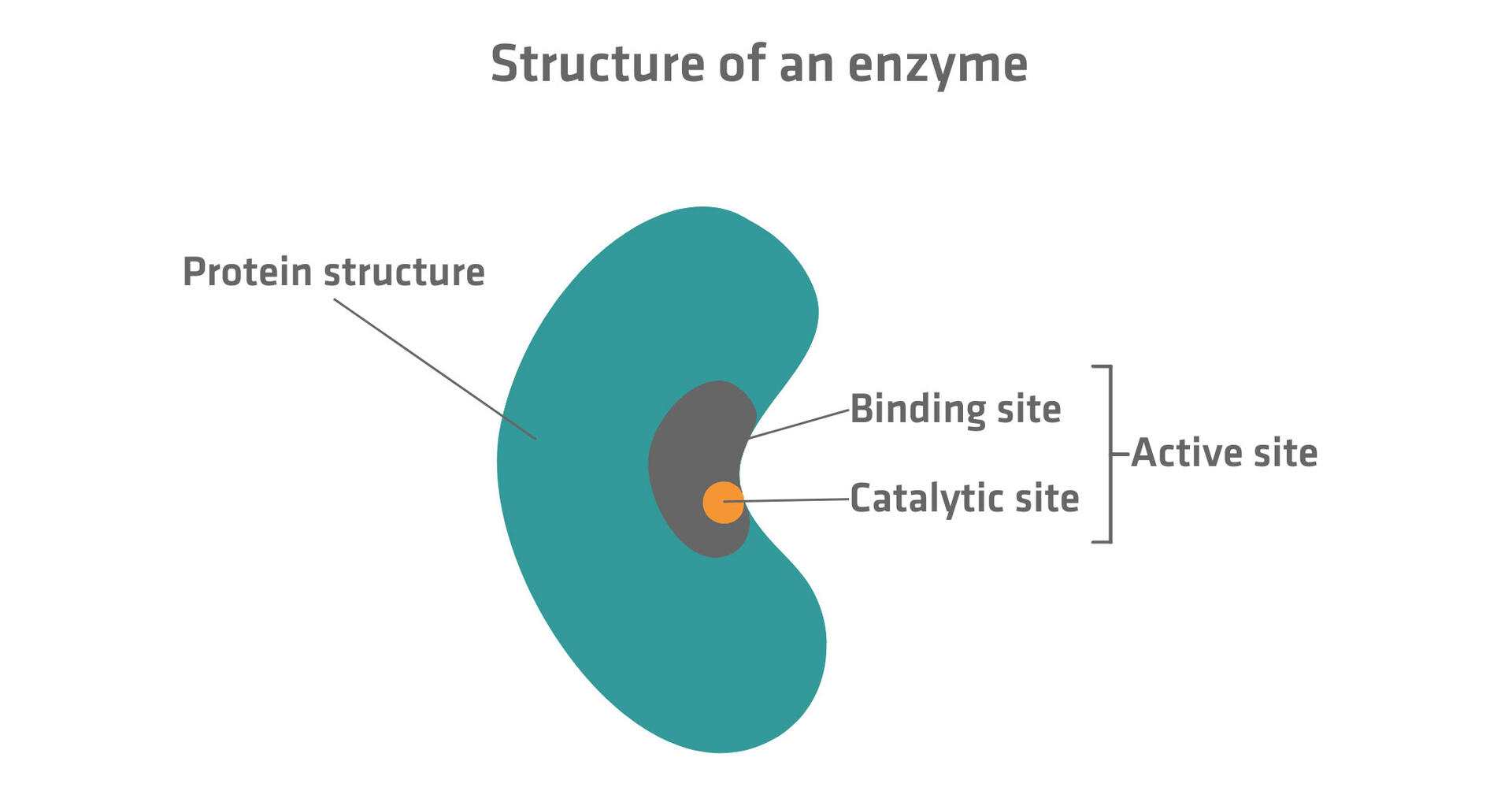
- Active site: this is a small portion of the enzyme that can reversibly bind and catalyze substrates. It looks like a groove or pocket, and consists of binding and catalytic sites.
- Binding site: the binding site contains specific amino acid residues that can form temporary bonds with substrate molecules and orient them.
- Catalytic site: the catalytic site is where the breaking and forming of bonds needed to transform the substrate molecule into one or several product molecules takes place.
This structure applies to most enzymes, but there are different types of enzymes called ribozymes that don't consist of proteins but of RNA molecules or RNA-protein complexes instead.4
Lock and key model vs induced fit model
Now that we understand how enzymes are structured, let's have a look at what happens during the different steps of the enzyme catalysis process. The first step is the reversible binding of the enzyme and the substrate (E + S → ES). For the substrate to bind to the enzyme, it must fit snuggly into the active site.
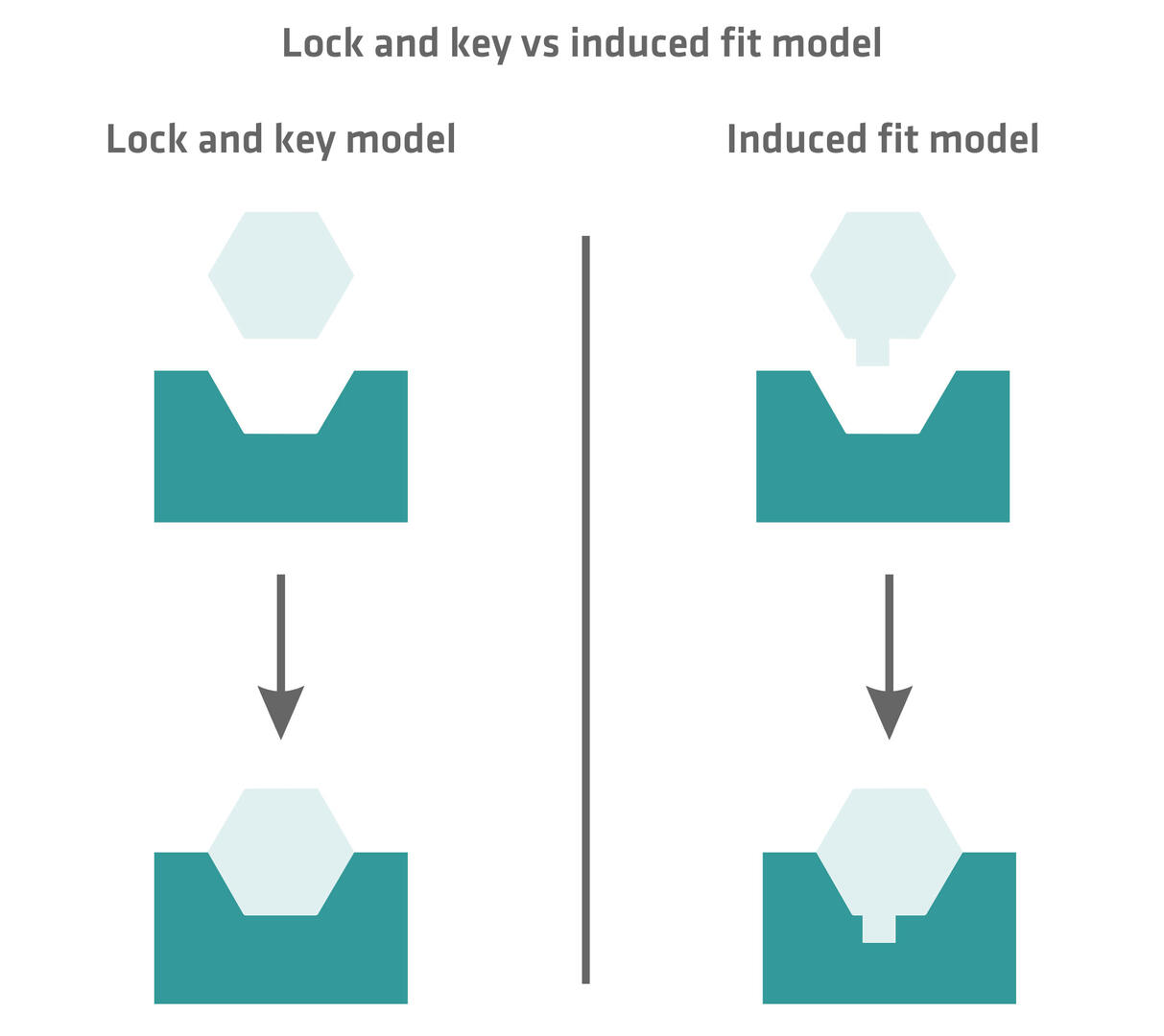
There are two models explaining the substrate binding process: the lock and key model and the induced fit model. The lock and key model – also called the Fisher model – assumes that the substrate fits precisely and perfectly into the active site due to their exact complementary shapes. According to the induced fit model – also referred to as the Koshland model – the active site is not exactly complementary to the substrate, but conforms to the structure of the substrate after they bind. This process is often compared to a glove (enzyme) changing its form when a hand (substrate) is inserted into the glove.
Catalysis mechanisms
Once the enzyme-substrate complex is formed, the chemical reaction converting the substrate into one or several products can take place (ES → EP). This transformation could also happen without enzymes, but it would take much longer – in many cases too long to sustain life – as the enzyme speeds up the reaction by lowering the activation energy.
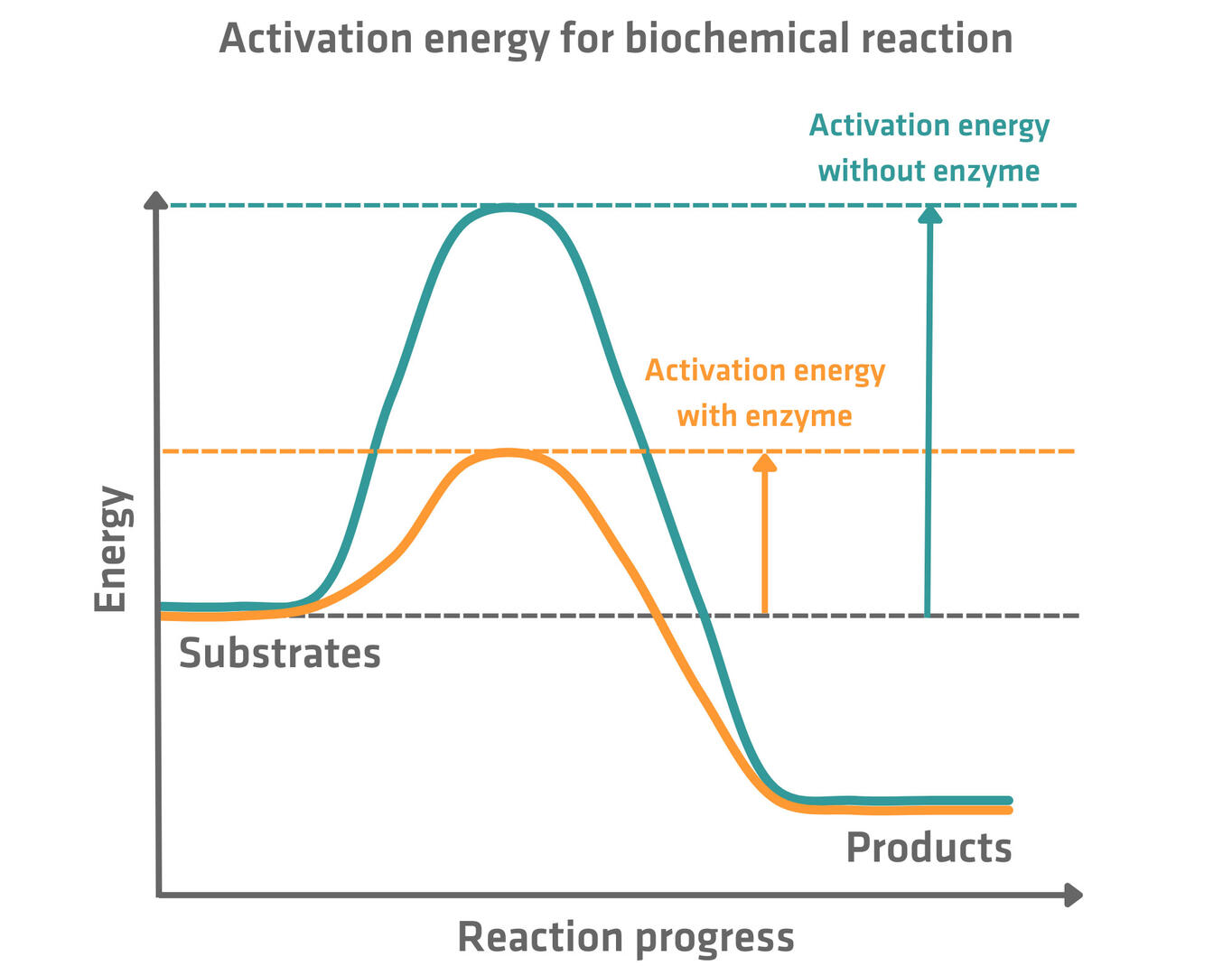
So, what happens within the active site of enzymes to lower the activation energy? There are several mechanisms that enzymes use to achieve this, and most enzymes in fact typically use a variety of different approaches. The four main mechanisms are:5,6
- Covalent catalysis: some enzymes lower the activation energy by temporarily forming a covalent bond with the substrate. Establishing a covalent bond modifies the electronic structure of the substrate and can enhance its reactivity.
- Catalysis by proximity and orientation: reactants must collide in a specific way for a biochemical reaction to happen. Enzymes can reduce the energy barrier by bringing reactants or substrates closer together for optimal orientation.
- Acid-base catalysis: the active sites of enzymes using acid-base catalysis contain specific residues that can accept or donate hydrogen ions. By transferring H ions, these residues can activate nucleophile and electrophile groups needed in the reaction, or stabilize charged groups and the transition state.
- Metal-ion catalysis: many enzymes use metal ions to ensure that reactions take place at reasonably high rates. Metal atoms can lose electrons and gain a positive charge, so they can facilitate interactions with different types of molecules in the active site. For example, a positive charge can help to form a strong nucleophile needed in the reaction, or hold and properly orient the substrate inside the active site.
If you'd like to learn more about these mechanisms, please refer to this video.
Reaction rate
After transforming the substrate into one or several products, enzymes release them (EP → E + P) and can bind to another substrate to start the enzyme catalysis process (E + S → ES) again.
The reaction rate of enzymes is affected by several factors briefly outlined below.
Temperature
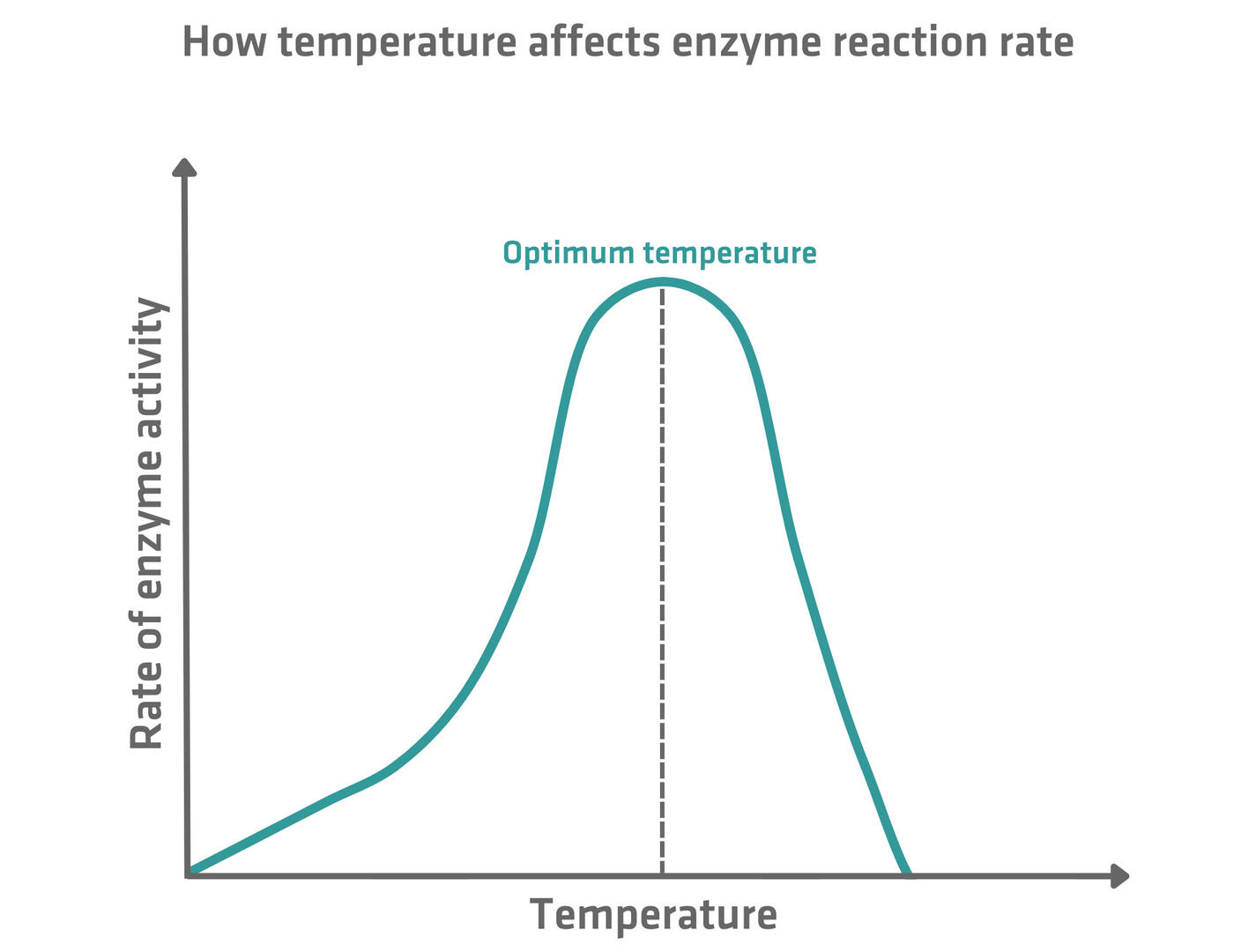
The rate of the enzyme catalysis process generally increases with temperature, reaching a peak at the enzyme's optimum temperature. Beyond this point, further temperature increases lead to a sharp decline in enzyme activity due to denaturation, where the enzyme's active site changes shape and loses functionality.7 The optimum temperature for an enzyme depends on where it normally works. Enzymes in human cells, for example, usually show the highest catalytic activity at around 37.5 °C.8
pH
Just as every enzyme has an optimum temperature, it also has an optimum pH value that varies based on its usual environment. Stomach enzymes, for example, have an optimum pH of about 2, whereas small intestine enzymes have an optimum pH of about 7.5.7
Salinity
The salinity of the environment affects the reaction rate, and each enzyme has an optimum salt concentration. Bonds and the 3D shape of enzymes are disrupted if the salt concentration is too low or too high, preventing substrate molecules from binding to the active site.9
Substrate concentration
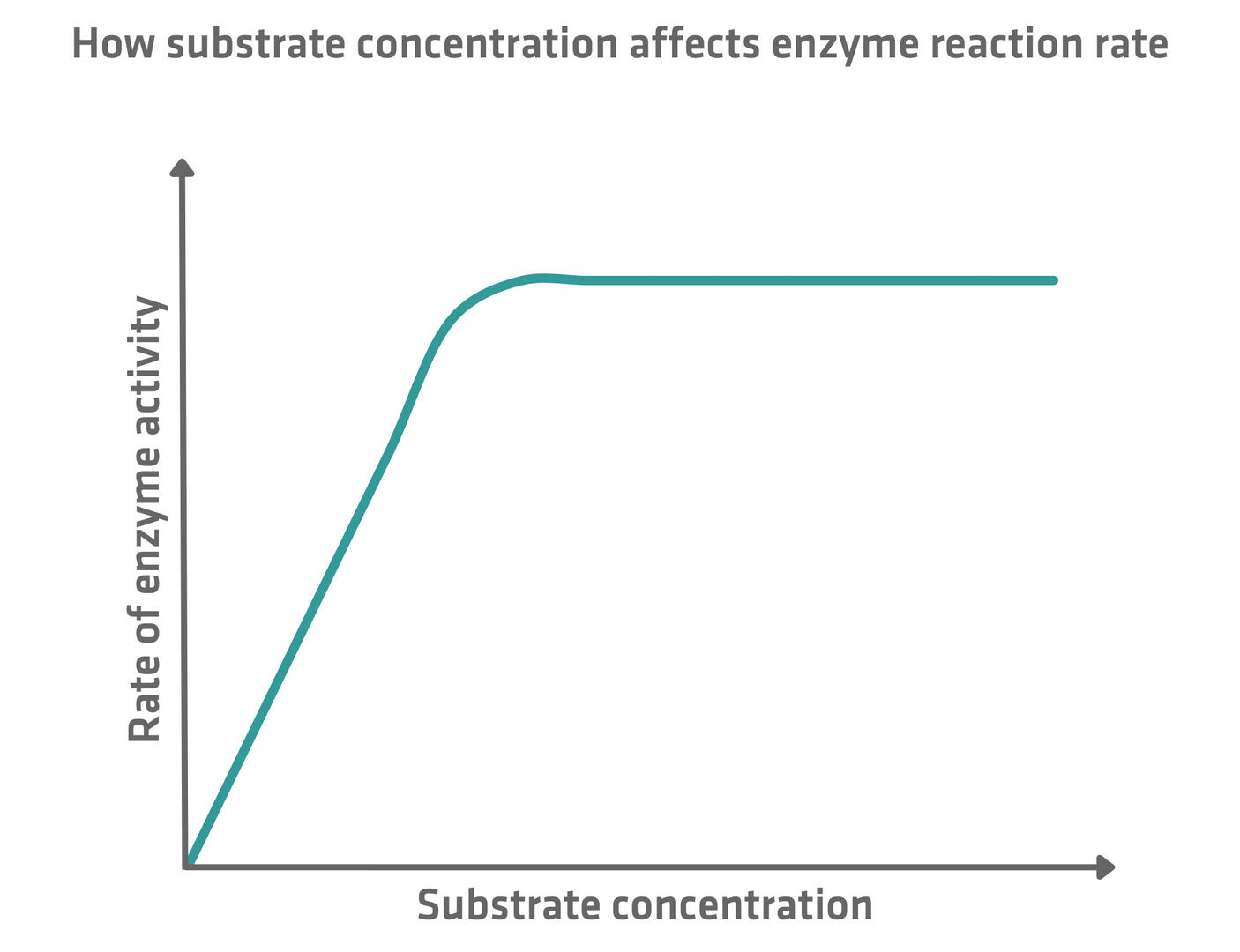
The reaction rate of enzymes depends on substrate concentration. Enzymatic activity rises as the concentration of substrate increases, because there are more opportunities for substrate molecules to bind to enzyme active sites. However, this increase in activity doesn't continue indefinitely. A saturation point is reached when all enzyme active sites are occupied. Beyond this optimum substrate concentration, the reaction rate remains constant, as there are no additional enzymes available to process the excess substrate.7
Co-factors and co-enzymes
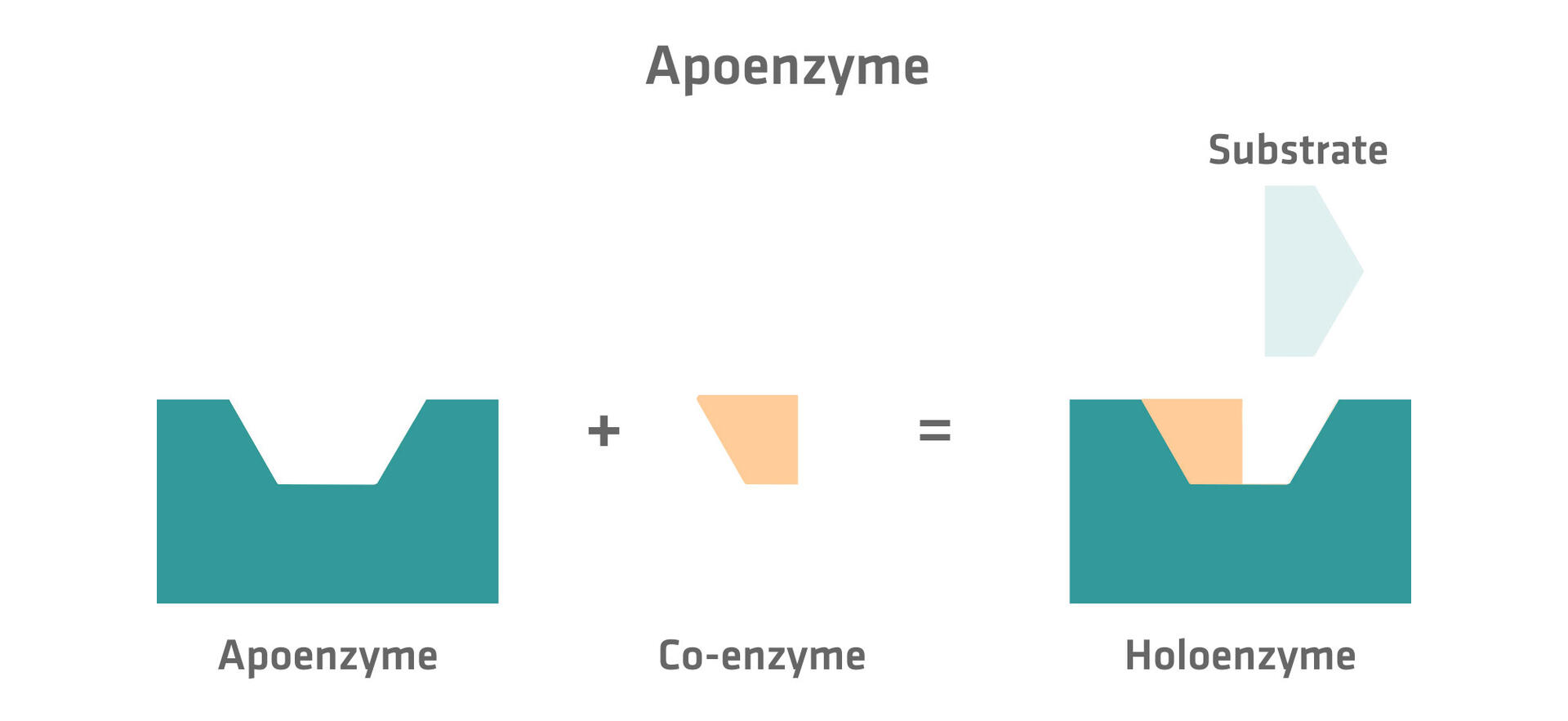
Co-factors and co-enzymes influence the reaction rates of apoenzymes. Apoenzymes are enzymes that are inactive until they are bound to a co-factor or co-enzyme. This catalytically active complex of the apoenzyme with the co-factor or the co-enzyme is called a holoenzyme.3
Co-factors and co-enzymes are both non-polypeptide molecules that are not part of the apoenzyme, but are needed to catalyze a reaction. Inorganic molecules are called co-factors and organic molecules co-enzymes. Examples of co-factors include ions like zinc or iron ions. Please note that the metal-ions used in metal-ion catalysis explained further above are co-factors. Examples of co-enzymes are vitamins or vitamin-derived molecules.10
Many co-factors and co-enzymes attach near the substrate binding site to facilitate the binding of the substrate to the enzyme.11
Enzyme inhibition
Enzymes can start the reaction catalysis process over and over again after transforming a substrate to a product, but biological systems such as human cells need the ability to stop or slow down the production of certain products when their concentration is high enough. That's where inhibitors regulating enzyme activity come into play. Moreover, certain drugs and toxins can also act as inhibitors.
There are two types of inhibition: reversible and irreversible inhibition.
Reversible inhibition
Reversible inhibition means that the inhibitors bind relatively weakly to enzymes so that they can easily dissociate again when environmental conditions change. Reversible inhibition can be further subdivided into competitive, uncompetitive and non-competitive.12
Competitive inhibition
Competitive inhibitors resemble the substrate and bind to the active site of enzymes.12
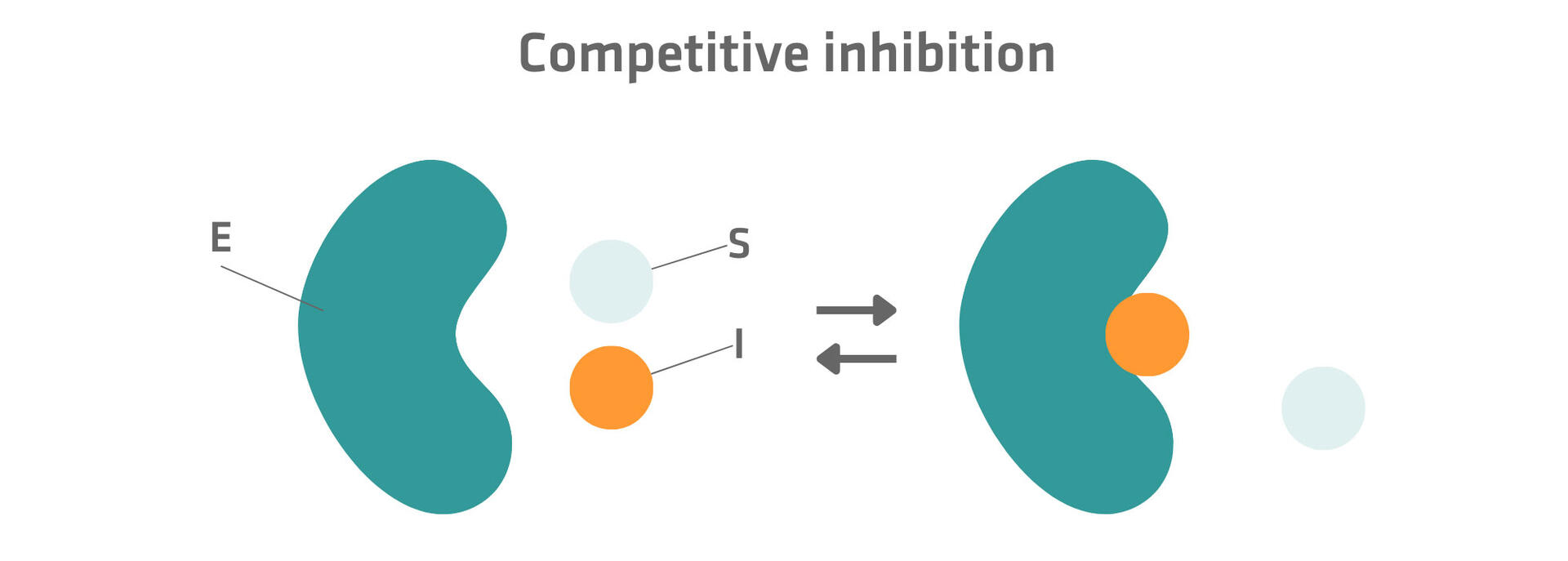
Competitive inhibitors typically have a higher affinity for the active site than substrates, and are more likely to occupy the active site. However, if the substrate concentration is increased, the substrate molecules can outcompete the inhibitor and remove the competitive inhibition effect.12
Please note that not all competitive inhibitors bind to the active site. Some of them can also bind to an additional binding site – a so-called allosteric site – and thereby change the composition of the active site so that the substrate can no longer occupy it. As only the substrate or the inhibitor can bind to the enzyme, this is also a type of competitive inhibition and called allosteric competitive inhibition.13
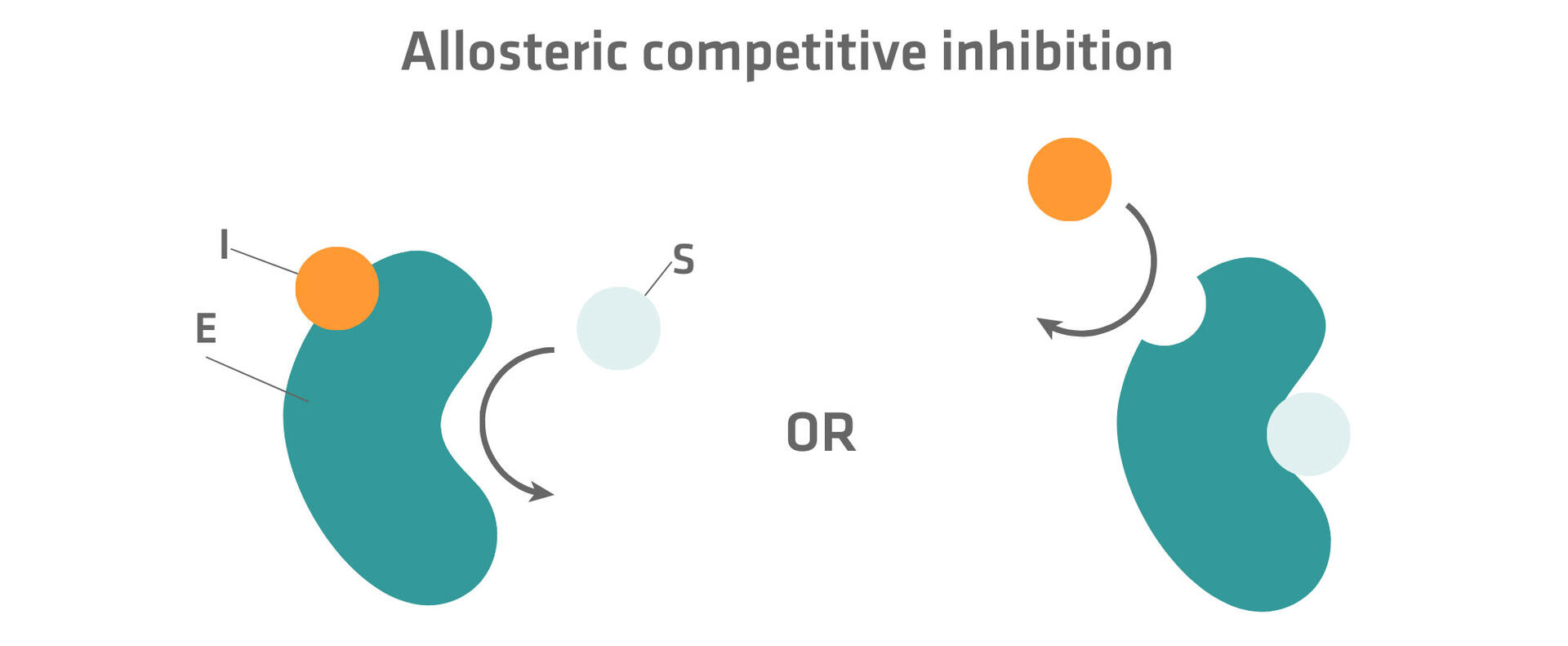
Uncompetitive inhibition
In some cases, the binding of the substrate to the active site changes the structure of the enzyme and creates an additional allosteric binding site. When an uncompetitive inhibitor binds to this allosteric site, the activity of the enzyme-substrate complex is blocked.12,14
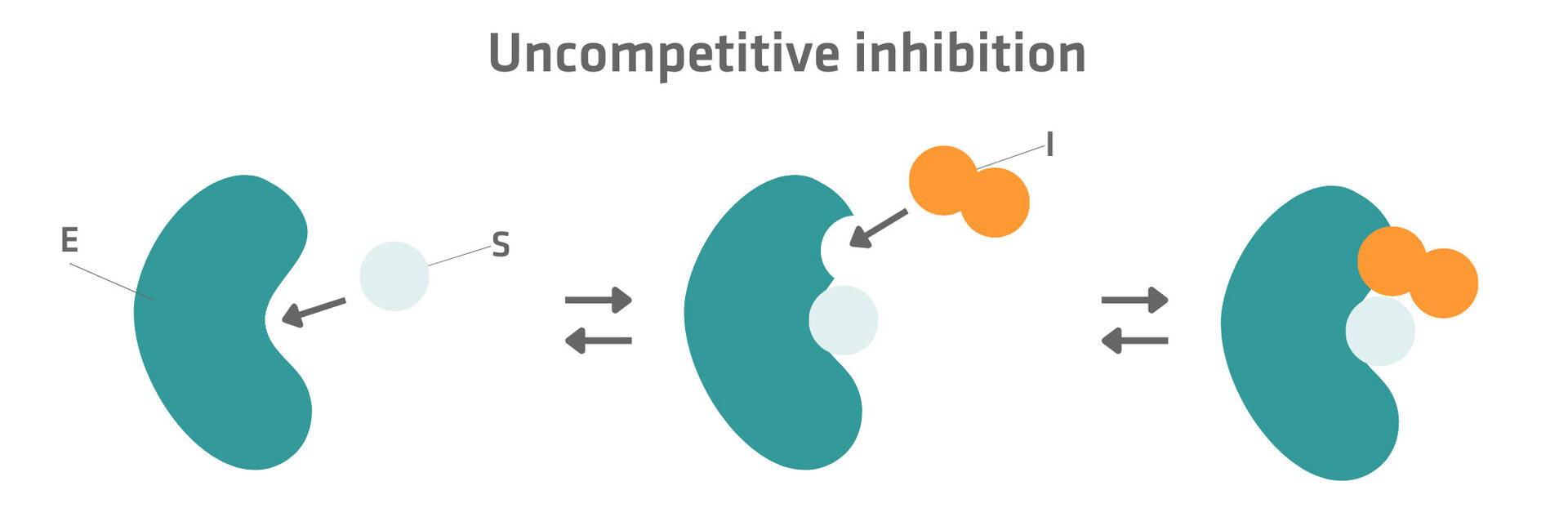
Please note that uncompetitive inhibition cannot be overcome by increasing substrate concentration.12,14
Non-competitive inhibition
Some enzymes have a permanent allosteric site that can bind to an inhibitor whether a substrate is bound or not. As substrates can also bind to the active site regardless of the presence of an inhibitor or not, several reaction pathways – as seen in the figure below – are possible. When an enzyme-substrate complex is formed, a product can be generated by the enzyme, whereas the enzyme-substrate-inhibitor complex remains inactive.14
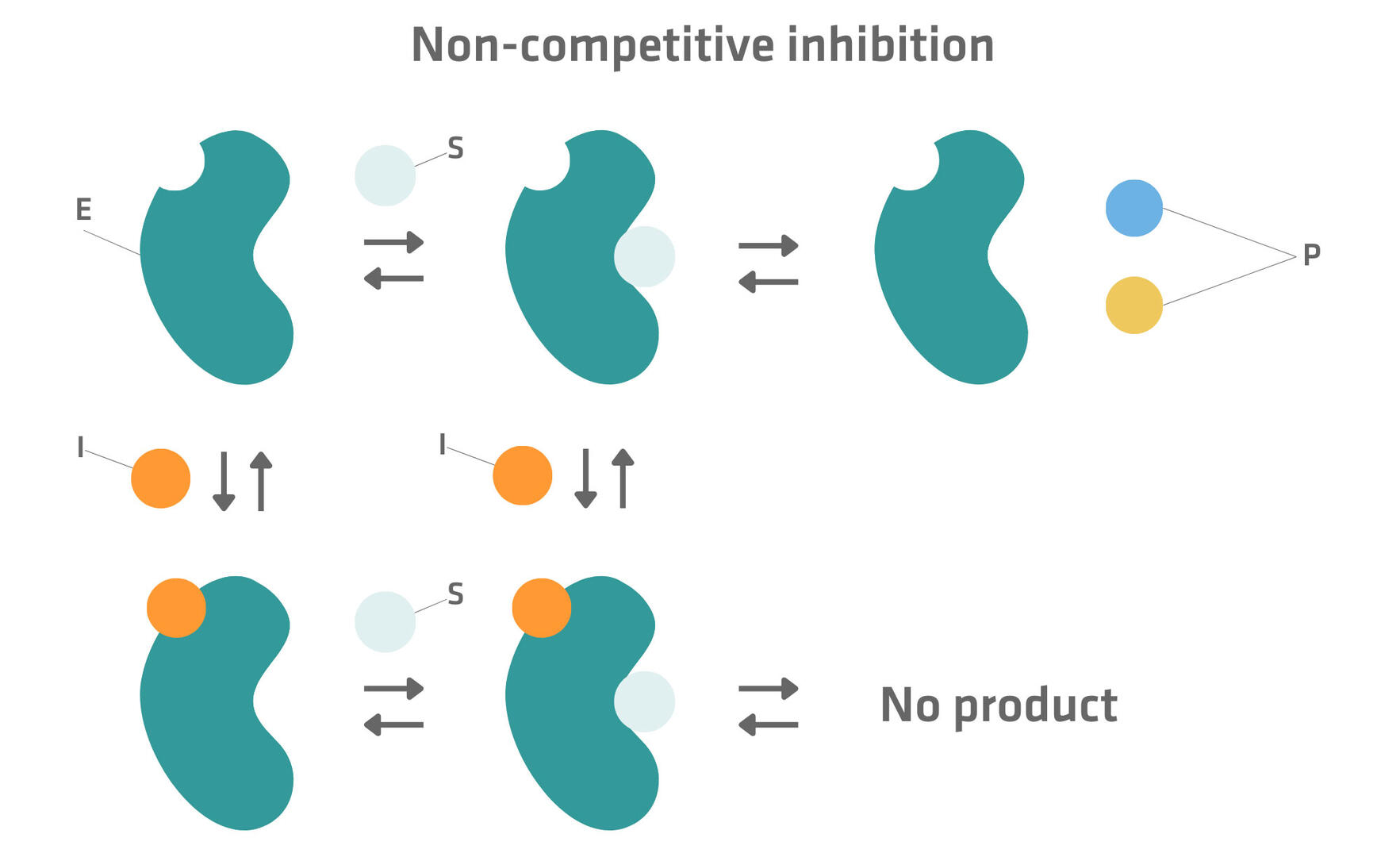
Some non-competitive inhibitors show an increased affinity for either the free enzyme or the enzyme-substrate complex. This type of non-competitive inhibition is then called mixed inhibition.15
Irreversible inhibition
Irreversible inhibitors block enzymatic activity by binding so tightly and strongly to enzymes that it's unlikely that they will ever dissociate again.12 For example, some pathogenic bacteria produce toxins that cause irreversible inhibition of protein synthesis in host cells.16 On the other hand, penicillin is a well-known irreversible inhibitor that prevents bacterial cells from forming cell walls, so that they eventually die.12
Feedback inhibition in metabolic pathways
As discussed in the example above, irreversible inhibition is mostly used to harm other organisms. Reversible inhibition, however, is a useful mechanism to inhibit metabolic pathways to ensure that cells always have the right product concentration at their disposal. For example, in multistep pathways, the end product often inhibits the first step of the reaction. This means that, when there is almost no end product available, the enzymes will work efficiently to fill up the stock, whereas a lot of excess end product molecules prevent the production of additional end products until the stock has been used up.17
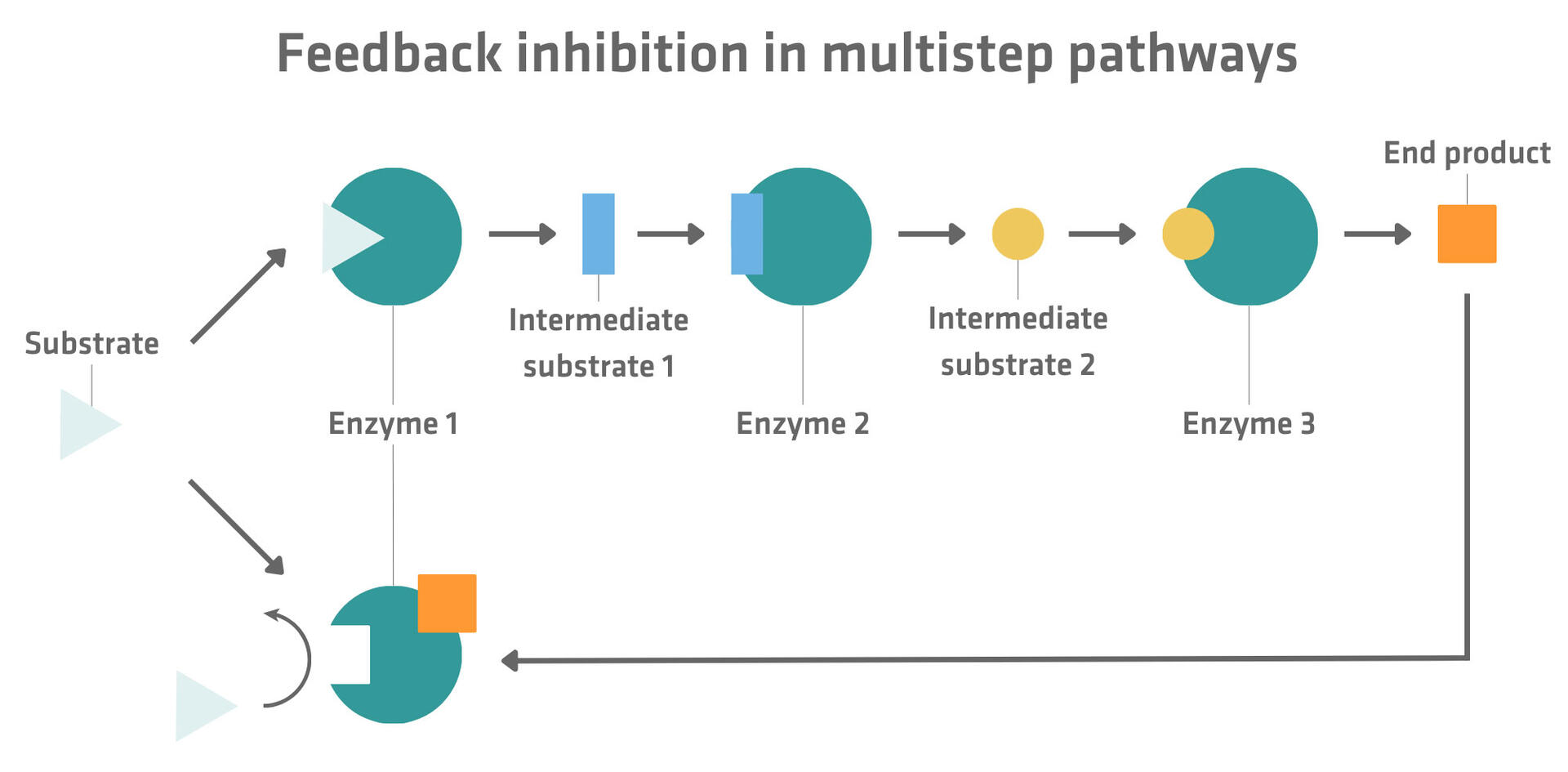
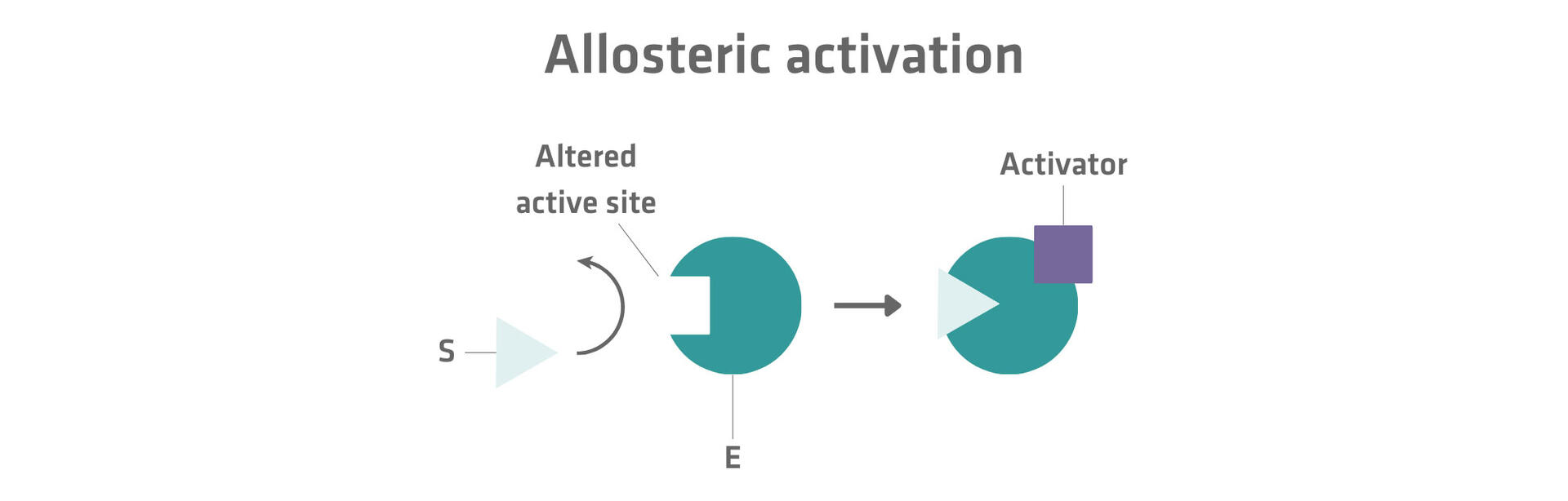
Please note that some products binding to enzymes don't inhibit but activate reactions. If they bind to an allosteric site, the conformation of the active site is changed in a way that the substrate can bind to it. This type of enzymatic regulation is called allosteric activation.17 For example, in some metabolic pathways, molecules that are produced when end products are used up by a cell can act as an allosteric activator for the first step in the chain of reactions producing the end product.18
Types of enzymes and their functions
Enzymes can be divided into seven different classes depending on the reaction that they catalyze. These classes have been defined by the International Union of Biochemistry and Molecular Biology (IUBMB) and are listed below.19
Class 1: oxidoreductases
Enzymes catalyzing the transfer of electrons from an electron donor (reductant) to an electron acceptor (oxidant) belong to the class of oxidoreductases.20
A– + B → A + B–
Class 2: transferases
Transferases are enzymes that transfer specific functional groups from one molecule (donor) to another (acceptor).21
AX + B → A + BX
Class 3: hydrolases
Hydrolases divide a large molecule into two smaller ones by using water to break chemical bonds.22
AB + H2O → A-H + B-OH
Class 4: lyases
Lyases catalyze the dissociation of a molecule into two distinct molecules without using water or oxidational reduction like hydrolases and oxidoreductases.23
A → B + C
Class 5: isomerases
Isomerases are involved in reactions where a molecule is being converted into one of its isomers.23
A → B
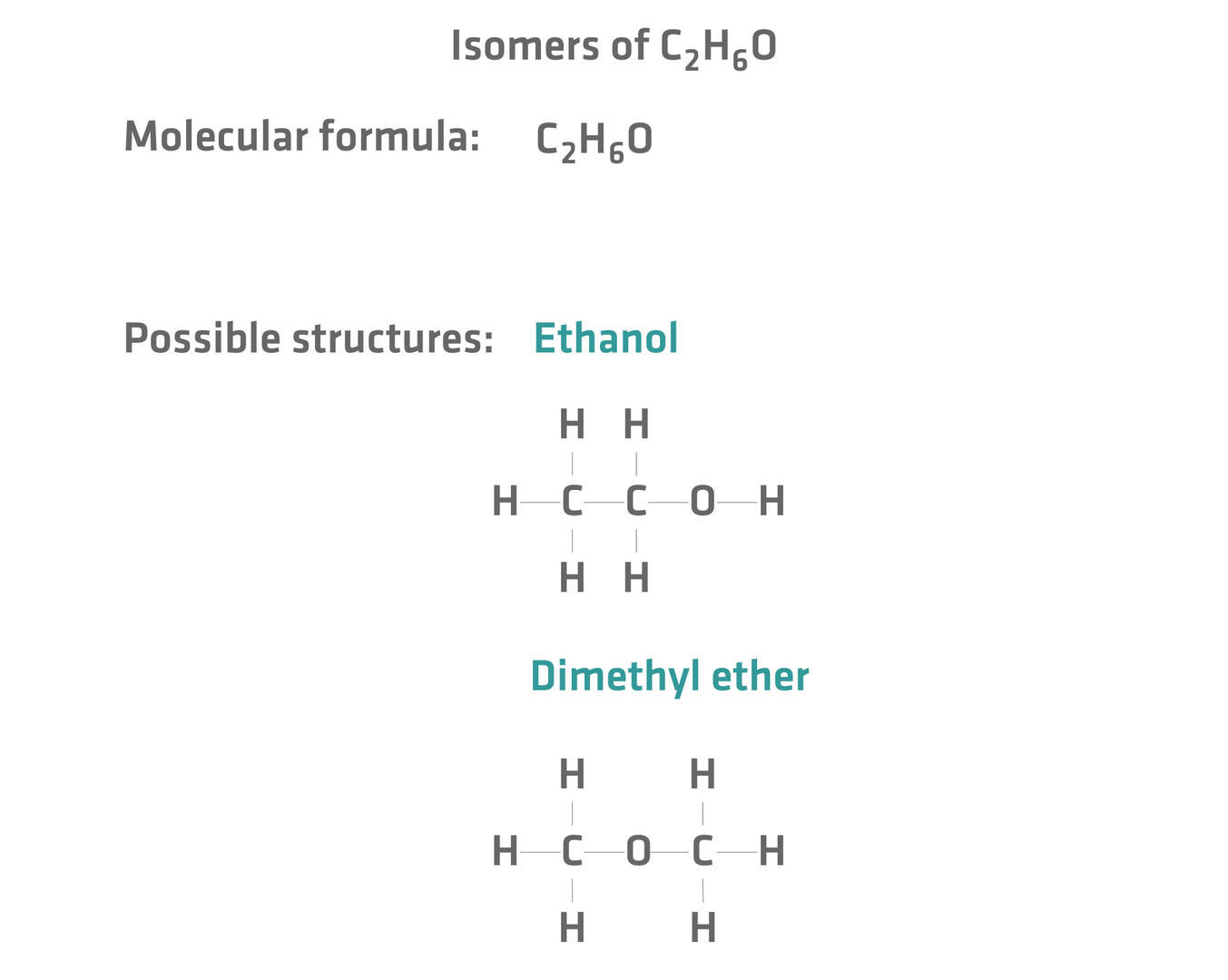
Isomers are compounds that have the same chemical formula but a different arrangement of atoms and consequently different properties.24
Class 6: ligases
Ligases join two molecules. This is usually coupled with the hydrolysis of a triphosphate, in most cases ATP.19,25,26
A + B + NTP → AB + NDP + P or AB + NMP + PP
NTP: Nucleoside triphosphate
NDP: Nucleoside diphosphate
NMP: Nucleoside monophosphate
Class 7: translocases
Translocases were only added to the list of enzyme classes in 2018. The definition of translocases is that they catalyze the movement of ions or molecules across membranes or their separation within membranes.26,27
AX + Bside 1 || = A + X + || Bside 2
As this formula can be a bit confusing, let's look at the example of H+-transporting two-sector ATPase. This enzyme converts ATP into ADP and phosphate, while transporting H+ to the other side of the membrane:28
ATP + H2O + 4 H+side 1 = ADP + phosphate + 4 H+side 2
Common applications of enzymes in the lab
As seen above, enzymes generally catalyze biochemical reactions within cells of living organisms. However, there are countless applications for enzymes in different industries, and enzymatic reactions form a part of several lab workflows. Here are a few examples:
- Polymerase chain reaction (PCR): PCR is used to copy a DNA sequence of interest millions or even billions of times. The enzyme making this possible is a transferase called Taq-polymerase and was isolated from the bacterium Thermus aquaticus discovered in a hot spring of Yellowstone National Park in 1969. To learn more about Taq-polymerase, please refer to our article 'How does PCR work and what are its 3 steps?'.
- Next generation sequencing (NGS): NGS enables us to determine the order of nucleotides in a DNA segment. While it took scientists years to sequence the entire human genome for the first time, this can now be accomplished within 48 hours thanks to NGS. Illumina and PacBio sequencing are two common NGS methods that use enzymatic reactions to ligate nucleic acid sequences to the sample DNA that is subsequently read by Taq-polymerase. Read our article 'DNA sequencing methods: from Sanger to NGS' to learn more.
- DNA fingerprinting: DNA fingerprinting can be used to determine an organism's genetic make-up and compare it to others. There are several DNA fingerprinting methods, including terminal restriction fragment length polymorphism (RFLP), which is commonly used to characterize microbial communities in environmental microbiology studies. DNA fragments extracted from a sample are amplified using Taq-polymerase and subsequently digested by restriction enzymes before being separated by size. Restriction enzymes are also called restriction endonucleases and belong to the enzyme class of hydrolases. Check out our article 'DNA fingerprinting: a powerful tool in forensics and beyond' to learn more about DNA fingerprinting and RFLP.
Conclusion
This guide provides a thorough introduction to the world of enzymes, highlighting their crucial role as biological catalysts. By exploring their structure, mechanisms of action, and the factors influencing their activity, we've gained insights into how enzymes facilitate and regulate biochemical reactions essential for life.
We hope that this article has provided a comprehensive overview to equip you with everything you need to know about enzymes. Leave a comment in the section below if you have additional questions and don't forget to subscribe to our newsletter.