Glossary
Find definitions for common laboratory, liquid handling and life sciences terms in our comprehensive glossary.
0 - 9
21 CFR Part 11
Part 11 of Title 21 of the Code of Federal Regulations (21 CFR Part 11) from the US Food and Drug Administration (FDA) establishes regulations on electronic records and electronic signatures. These regulations are highly relevant to GCP, GLP and GMP regulated laboratories, and provide a definition of what it takes for the FDA to trust electronic records when they are submitted to, or viewed by, FDA reviewers and investigators.
3D cell culture
3D cell culture refers to the process of growing cells in three dimensional cultures that mimic the in vivo environment. This allows the study of complex interactions between cells, as well as between cells and the extracellular matrix (ECM). In contrast to 2D cultures, where cells are mostly exposed to the media, all cells are exposed either to other cells or to the ECM in 3D cell cultures.
A
Accuracy
See pipette accuracy.
Adjustable tip spacing pipette
See spacer pipette.
ADME
In drug development, the abbreviation ADME stands for absorption, distribution, metabolism and excretion. The characterization of ADME properties helps to explore and explain how pharmacokinetic processes happen, and provide safety considerations of a new drug.
Affinity chromatography
Affinity purification
Affinity purification, also called affinity chromatography, is a chromatographic method used to separate a specific molecule from a complex mixture. It is based on highly specific biological interactions between two molecules, such as interactions between enzyme and substrate, receptor and ligand, or antibody and antigen. These interactions, which are typically reversible, are used for purification by placing one of the interacting molecules, referred to as affinity ligand, onto a solid matrix to create a stationary phase, while the target molecule is in the mobile phase.
Air displacement pipette
Air displacement pipettes have an air cushion between the piston and the sample. To aspirate liquid, the piston first descends, pushing out a volume of air corresponding to the required volume of liquid, before moving up again and creating a partial vacuum that will be filled by the liquid.
Amplicon
An amplicon is a DNA or RNA sequence that is the source and/or product of amplification or replication events. Amplicons are naturally produced through gene duplication and can be artificially generated through methods such as polymerase chain reactions.
Antibody
Antibodies (also called immunoglobulins) are protein molecules used by the immune system to recognize and neutralize foreign objects – such as pathogenic bacteria and viruses – and cancer cells. Antibodies bind to a specific antigen on the outside of a foreign object or cancer cell to neutralize it or to tag it for attack by other parts of the immune system. In patients with autoimmune diseases, antibodies sometimes mistakenly recognize normal cellular proteins as antigens.
Antigen
Antigens are molecular structures – usually composed of proteins, peptides and/or polysaccharides – that trigger an immune response. They can be divided into exogenous, self and tumor antigens:
- Exogenous antigens can be found on the outside of foreign objects entering the body. For example, any portion of a pathogenic bacteria or virus, such as a surface protein, can serve as an antigen.
- Self antigens are normal cellular proteins that are mistakenly attacked by the immune system in patients with autoimmune diseases.
- Tumor antigens are expressed on the cell surface of cancer cells. However, they are often not detected by the immune system, as most cancer cells can escape immune system-mediated elimination.
Aqueous solution
In an aqueous solution, one or more substances (solutes) are dissolved in water (solvent). Water dissolves only hydrophilic substances such as acids, bases and many salts. In a chemical equation, the symbol (aq) indicates that a certain substance is dissolved in water, e.g. NaCl(s) → Na+(aq) + Cl-(aq)
Automated liquid handling systems
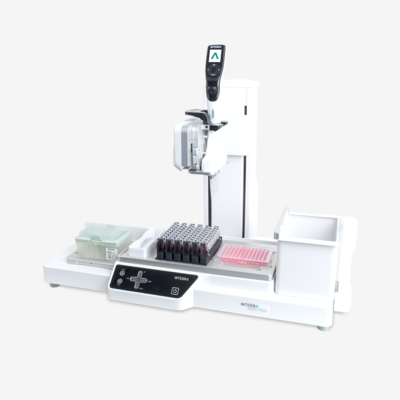
Automated liquid handling systems, also called liquid handling platforms or pipetting robots, release researchers from repetitive and laborious tasks, reduce processing time, decrease sample contamination and increase reproducibility. Researchers can choose between investing in a large system for large scale processing, or a compact one that fits into a laminar flow hood. Benchtop versions are also available, without compromising system complexity or handling capacity.
Automated pipette
Automated pipettes are simple, benchtop automated liquid handling workstations that avoid manual labor for repeated pipetting, offering improved throughput and reproducibility.
B
Bacterial conjugation
Bacterial conjugation is a process of horizontal gene transfer by which genetic material is transferred from a donor bacterium to a recipient bacterium through cell-to-cell contact. Donor bacteria that contain a DNA sequence called the fertility factor can create a pilus that attaches to the recipient. Once in contact, the donor can transfer genetic material, commonly a plasmid.
Bacterial culture
Bacterial culture refers to the process of growing bacteria in or on a medium in a lab environment. The culturing parameters vary between the different bacterial species. One of the most important factors is oxygen. Depending on whether your bacteria need oxygen or not, you have to grow the bacterial cells in an aerobic or anaerobic culture.
Bacterial transduction
Bacterial transduction is a process of horizontal gene transfer by which bacterial DNA is transferred from one bacterium to another by a bacterial virus (also called bacteriophage). First, a bacteriophage attaches to a bacterial cell and injects its genetic material. When new bacteriophages are generated inside that bacterium, bacterial DNA is then sometimes accidentally incorporated into the bacteriophages' DNA. As soon as the bacteriophages emerge from the host, attach to another bacterial cell, and inject their genetic material, the bacterial DNA is transferred.
Bacterial transformation
Bacterial transformation is a process of horizontal gene transfer by which some bacteria take up foreign genetic material from the environment and integrate it into their chromosome. It does not require a living donor cell, but only the presence of persistent DNA in the environment. Bacteria that can take up free, extracellular genetic material are called competent cells and undergo transformation to respond to changes in the environment, e.g. carbon starvation.
Biochemistry
Biochemistry explores the chemical processes within and related to living organisms, focusing on processes happening at a molecular level. This covers a range of scientific disciplines, including genetics, microbiology, forensics and medicine.
Biological liquid waste disposal
Liquid laboratory waste may contain chemical, radioactive, human/animal or other biological material that could potentially harm the environment or population. The correct disposal of biological liquid waste includes waste segregation, total containment, labeling and deactivation.
Biosafety cabinet (BSC)
Biosafety cabinets (or biological safety cabinets) are enclosed, ventilated workspaces under negative pressure, equipped with high efficiency particulate air (HEPA) filters. They are used in laboratories working with biologically hazardous materials to protect personnel, the environment and – depending on the type of biosafety cabinet – the products in the cabinet from aerosols and contamination.
Blow-in
In air displacement pipettes, the piston moves back into its initial position after a blow-out. This causes a slight intake of air and is called blow-in.
Blow-out
Air displacement pipettes expel extra air after a dispense to discharge any residual liquid from the pipette tip. This process is called blow-out.
Buffer solution
A buffer is an aqueous solution consisting either of a mixture of a weak acid and its conjugate base, or a weak base and its conjugate acid, which is resistant to changes in pH. Buffers are used to maintain a stable pH in a solution, as they can neutralize small quantities of additional acid or base.
C
Calibration
See pipette calibration.
Cell banking
Cell banking refers to the process of storing multiple batches of a specific cell type for later use, to avoid factors such as random mutations or transcription errors that compromise reproducibility. The first step is establishing a Master Cell Bank (MBC) by growing a selection of the cell type of interest in a culture, and cryopreserving it in multiple containers. The MBC batches are thawed and used to prepare Working Cell Banks (WBC) later on.
Cell culture
Cell culture refers to the process of growing cells under controlled conditions outside their natural environment. It is an in vitro tool that aids the understanding of cell biology and the mechanisms of diseases. It also plays a role in drug discovery for applications such as drug toxicity testing and pharmacokinetic/pharmacodynamic studies, as well as in personalized medicine.
Cell culture contamination
Cell culture contamination can impact cell viability and, if undetected, the reliability of results by leading to inaccurate conclusions. The two main sources of cell culture contamination are chemical and biological contaminants. Examples of chemical contaminants include impurities in media, sera and water, endotoxins, plasticizers, and detergents, whereas biological contamination is mostly due to bacteria, molds, yeasts, viruses, mycoplasma or the cross-contamination by other cell lines.
Cell culture medium
A cell culture medium, also known as growth medium, is a liquid or gel created to support cellular growth in an artificial environment. It is comprised of a combination of compounds and nutrients, such as amino acids, vitamins, carbohydrates, inorganic salts, basic and trace elements, serum, hormones, buffering systems, supplements and antibiotics.
Cell culture medium change
To keep cells in cell cultures healthy, the medium needs to be changed regularly to provide fresh nutrients. In the case of adherent cell cultures, the media can be aspirated and replaced. In non-adherent cell cultures, the cells need to be centrifuged and resuspended in fresh medium.
Cell line development
A cell line is a permanently established cell culture that will proliferate indefinitely given appropriate fresh medium and space, either because of a random mutation or deliberate modification. For cell line development, researchers have access to several non-genome-editing methods – including single cell cloning, cell line adaption, differentiation and immortalization – or gene editing technologies such as CRISPR.
Cell lysis
Cell lysis refers to the breaking down of the cell membrane. Cell lysis can occur naturally, for example through a viral infection or osmosis, or be purposely induced to study the cell's components. Cell lysis techniques can either be mechanical (e.g. bead mill), physical (e.g. heating), chemical (e.g. detergent) or enzymatic.
Cell seeding
The procedure of spreading a defined amount (volume or cell number) of a cell suspension into a new flask or plate is called cell seeding. If you are working with adherent cells, you need to use trypsin to detach them from the surface of the current cell culture vessel before seeding.
Cell staining
Cell staining is a technique used to visualize cells and cell components under a microscope. Positive stains allow you to highlight certain cell components – such as the nucleus, the cell wall or the entire cell – whereas negative stains are absorbed by the background outlining the cells.
Cell viability assay
Cell viability assays are used to determine the overall health of cells by measuring the proportion of live, healthy cells within a population.
Centrifugation
During centrifugation, the contents of a sample tube are spun at a defined speed to separate the contents based on their relative densities.
Centrifuge tube
Centrifuge tubes are made especially to hold liquids during centrifugation. They usually have conical bottoms, and are available in a range of different materials and sizes. Tubes with a capacity of 2 ml or less are often referred to as microcentrifuge tubes.
Chemical compound
A chemical compound consists of molecules formed from at least two different elements. The chemical formula of the compound, e.g. H2O, indicates what elements make up the compound, and how many atoms of each element are contained in one molecule.
Cherry picking
See hit picking.
Coefficient of variation (CV)
The coefficient of variation (CV) is defined as the extent of variability in relation to the mean. If a pipette has a low CV, it means that it dispenses volumes close to the set volume, leading to precise results.
COVID-19
COVID-19 is a highly contagious respiratory disease caused by the SARS-CoV-2 virus. The most common signs and symptoms of COVID-19 are fever, cough and breathing difficulties. Fatigue, muscle pain, chills, headache, sore throat, runny nose, nausea or vomiting, diarrhea, and a loss of taste or smell may also occur. The signs and symptoms may be mild or severe and usually appear 2 to 14 days after exposure to the SARS-CoV-2 virus. Some people may not have any symptoms, but are still able to spread the virus.
CRISPR-Cas9
CRISPR-Cas9 is a genome editing technology used to change parts of the genome by removing, adding, or altering sections of DNA. It uses a pre-designed RNA sequence to guide the Cas9 enzyme to the location where the genome needs to be altered. The Cas9 enzyme then cleaves the DNA strands, and the cell's natural repair mechanism joins the two ends together by removing, adding, or replacing certain sections.
D
De novo assembly
De novo assembly is used to study organisms whose genomes have not previously been sequenced. It refers to the process of constructing an organism's genome from scratch, using only sequences of DNA obtained through sequencing, without the use of a reference genome.
Dead volume
The dead volume is the residual volume of liquid that cannot be recovered from a reagent reservoir, because it cannot be aspirated by the pipette or any other device.
Detergent
Detergents contain a hydrophobic tail and a hydrophilic head and can therefore disrupt or form hydrophobic-hydrophilic interactions among molecules in biological samples. These properties make them useful to lyse cells, solubilize proteins and lipids, control protein crystallization, prevent nonspecific binding in affinity purification and immunoassay procedures, and for gel electrophoresis.
Dimethyl sulfoxide (DMSO)
Dimethyl sulfoxide (DMSO) is a polar, aprotic organic solvent commonly used as cryoprotectant in cell culture, or to prevent the formation of secondary structures in DNA templates or primers during PCR.
Dispensing cassette
Dispensing cassettes are reagent dispenser accessories for accurate and rapid reagent addition into 6, 12, 24, 96, 384 and 1536 well plates. They usually consist of 8 tubes and are available in different bore sizes, although some manufacturers also offer 16 channel dispensing cassettes to make the filling of 384 and 1536 well plates more efficient.
DMPK
Drug metabolism and pharmacokinetics (DMPK) is a core discipline in drug development that considers the biotransformation and other pharmacokinetic properties of a drug to assess its safety. DMPK studies allow drug developers to experimentally evaluate intrinsic properties of a drug candidate to validate that it can and will be cleared from the body without producing harmful byproducts (metabolites), reaching dangerous exposure levels (toxicity), or causing adverse side effects.
DNA
Deoxyribonucleic acid (DNA) stores the hereditary material in all organisms. It consists of a double helix formed by two single strands made from the nucleobases (adenine, guanine, cytosine and thymine) and a sugar-phosphate (deoxyribose and a phosphate group) backbone.
DNA extraction
DNA microarray
A DNA microarray, sometimes called DNA chip, is a collection of microscopic DNA spots attached to a solid surface. Each spot contains thousands of copies of a specific, single-stranded DNA sequence, known as probes. DNA microarray experiments can be used to visualize gene expression or for genotyping assays.
For example, to determine whether an individual possesses a mutation for a particular disease, a DNA sample from the patient and a control sample are denatured, cut into fragments and commonly labeled with a fluorescent dye (red for patient's sample, green for control sample). Both samples are inserted into the microarray and allowed to hybridize to the single-stranded DNA on the chip. When scanning the microarray, yellow spots indicate DNA sequences shared by the patient and the control sample, whereas green spots represent the 'normal' sequence and red spots mutations.
DNA polymerase
A DNA polymerase is an enzyme using dNTPs to create complimentary copies of specific DNA sequences during PCR reactions. Taq-polymerase is typically used, which was originally isolated from the bacterium Thermus aquaticus, discovered in a hot spring in the Yellowstone National Park in the 1960s.
DNA purification
DNA sequencing
DNA sequencing is used to determine the order of nucleotides in a DNA segment, genome, or microbiome. The steps needed to obtain the order of nucleotides depend on the sequencing method chosen, with common techniques including Sanger, Illumina, PacBio and Nanopore sequencing.
DNA-encoded library (DEL)
A DNA-encoded library is a collection of chemical compounds or building blocks that are covalently attached to short DNA fragments. Similar to barcodes, the DNA fragments can be used to identify the compounds or blocks, and in some cases, they can also direct and control their chemical synthesis.
DNA-encoded libraries are used in drug discovery to accelerate early phase activities (see DNA-encoded library screening).
DNA-encoded library (DEL) screening
To screen a DNA-encoded library, the mix of compounds and their DNA tags is combined with a biological target, such as an enzyme. Compounds that do not bind to the target wash away. The target is then denatured, and the resulting batch of ‘hits’ is incubated with a fresh target to ensure that the best binders remain. After repeating this process a third time, the DNA tag on each compound is amplified and sequenced, to determine which compounds bound best to the target. These compounds are then resynthesized without the DNA tag, and tested with the target.
DNase
See nuclease.
dNTPs
Deoxyribonucleotide triphosphates (dNTPs) are the 'building blocks' of DNA. There are four different types of dNTPs: dATPs, dCTPs, dGTPs and dTTPs. During PCR reactions, dNTPs are needed to create complimentary copies of the DNA sequence of interest.
Drug screening
Drug screening is the process of identifying and optimizing potential drugs before selecting a compound for clinical trials. It can involve screening vast compound libraries for a particular biological activity in high throughput assays.
Dry touch off
See touch off.
E
E. coli
Escherichia coli (E. coli) is a Gram-negative, rod-shaped bacterium that is commonly found in the lower intestine of warm-blooded organisms. Most E. coli strains are harmless, but some serotypes are pathogenic and can cause serious food poisoning in humans and other species. While there is a great diversity of E. coli strains in the environment, only a few are used in the lab, mostly K-12. K-12 is safe to handle, easy to grow and divides every 20 minutes under optimal conditions.
ELISA
An ELISA test is a type of enzyme immunoassay (EIA). These highly specific and sensitive assays are used to detect concentrations as low as 0.01 nanograms of antigen or antibody per milliliter. When performing ELISAs, antigen-antibody complexes are immobilized to a solid surface. An enzyme is covalently attached to one of the molecules in the complex, and the subsequent addition of an enzyme-specific substrate results in a colored reaction product.
Endotoxin test
The endotoxin test is an in vitro assay used to detect bacterial endotoxins, a component of the cell wall of Gram-negative bacteria. This test is performed for medical devices with direct or indirect contact to the cardiovascular system, lymphatic system, or cerebrospinal fluid. Injectable pharmaceutical products must also be tested for bacterial endotoxins.
Epitope
The epitope (also called antigenic determinant) is the part of the antigen that triggers an immune response. If an epitope binds to a complementary antigen receptor on a B cell, the B cell will start to produce antibodies specifically targeting this epitope. Many antigens have several epitopes on their surface, which means that various antibodies targeting different epitopes can bind to an antigen at the same time.
F
Flow cytometry
Flow cytometry is a method for the qualitative and quantitative measurement of biological and physical properties of cells and other particles. Particles are suspended in a high-velocity fluid stream, and passed through a laser beam one at a time.
Fluorometry
Fluorometry is a technique used to identify and quantify analytes in a sample by adding fluorophores that bind to the analyte of interest to the sample, exciting them with a beam of UV light and detecting and measuring the emitted wavelength. By comparing the sample fluorescence to known standards, the analyte can be quantified.
Forward pipetting
Forward pipetting is the most commonly used pipetting technique to aspirate and dispense a certain volume of liquid with an air displacement pipette. It consists of the following steps:
- Push down the plunger to the first stop
- Immerse the pipette tip into the liquid and release the plunger
- Remove the pipette tip from the liquid
- Push the plunger down to the first stop to dispense the liquid
- Depress the plunger to the second stop to expel any residual liquid
- Remove the pipette tip from the vessel and release the plunger
G
Gel electrophoresis
Gel electrophoresis is a technique used to separate molecules like DNA, RNA or proteins based on their size and charge. Samples are loaded into wells at one end of a gel, and an electric current is applied. The molecules travel faster or slower through the gel depending on their charge and size, allowing them to be separated from one another.
Gene expression
Gene expression is the process by which information encoded in DNA is converted into a protein molecule. It consists of two steps, transcription and translation. During transcription, the information stored in DNA is converted into messenger RNA (mRNA), a sort of ‘instruction for making proteins’. The mRNA then leaves the cell nucleus, and the ribosome translates it into a protein, reading codon by codon and adding the corresponding amino acid for each sequence to the protein.
Gene isoform
Gene isoforms are different mRNA molecules produced by the same gene. These mRNA sequences often differ in their protein-coding capacity, expanding the functional diversity of the proteins a single gene can produce.
Genotyping
Genotyping is a molecular assay designed to detect the presence or absence of a specific gene variant (or variants, also called alleles) in an individual. It can be used, for example, to screen for mutations that can predispose individuals to certain diseases.
Good Laboratory Practice (GLP)
The OECD Principles of Good Laboratory Practice (GLP) are a quality control system covering the organizational process and the conditions under which non-clinical health and environmental studies are planned, performed, monitored, recorded, reported and retained or archived.
H
Helicase-dependent amplification (HDA)
Helicase-dependent amplification (HDA) is a technique used for isothermal DNA amplification. Unlike PCR, it does not require several thermal cycles, as a helicase that unwinds DNA strands eliminates the need for heat denaturation.
Hit picking
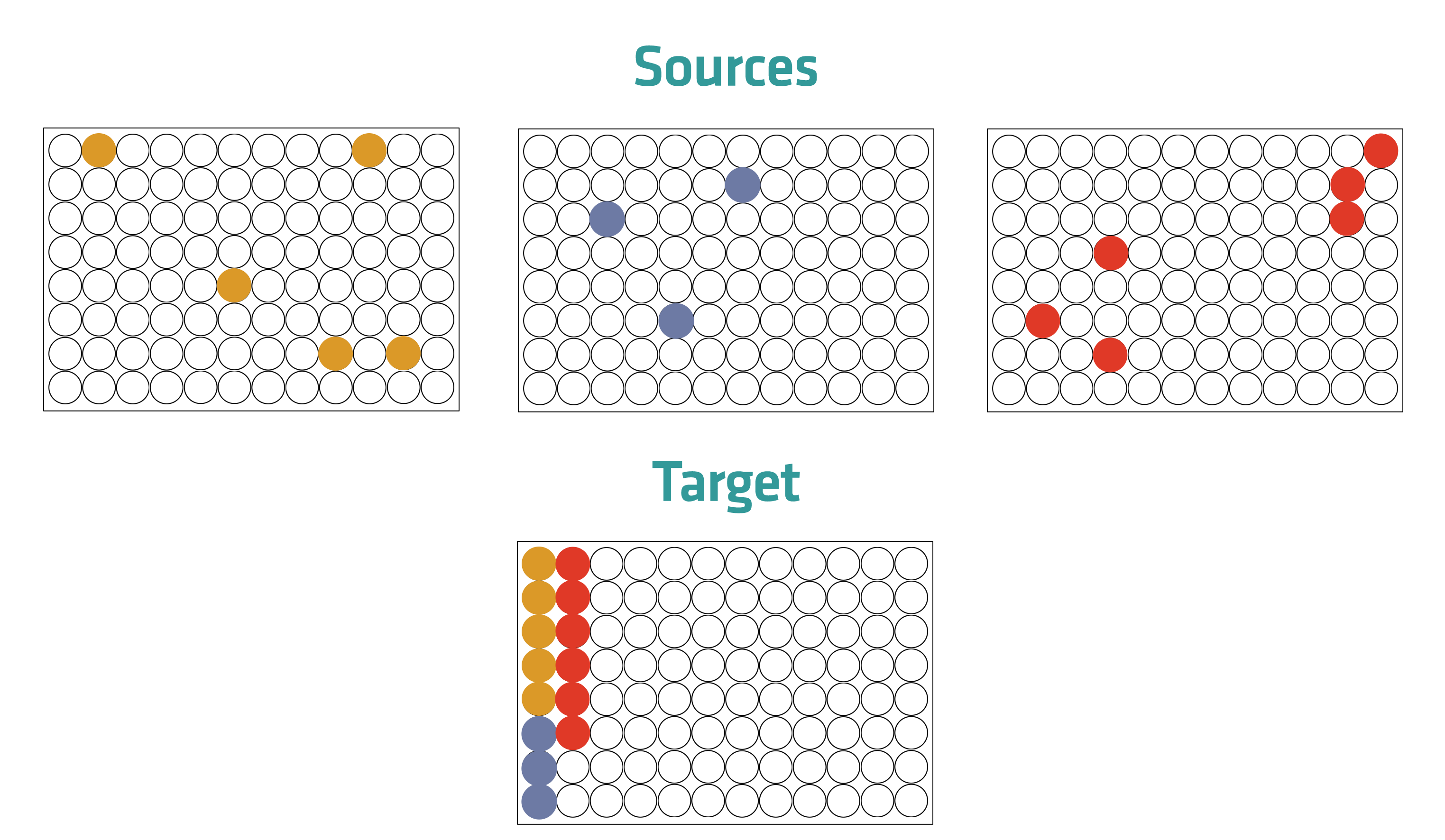
Hit picking (also called cherry picking) is the process of condensing ‘hits’ by transferring them from multiple sources to one destination, e.g. from multiple microplates to one microplate. Hits are samples, such as drug compounds, that have been identified as compatible or effective in a previous screening assay.
Homopolymer
A homopolymer is a repetition of identical bases – adenine (A), thymine (T), cytosine (C), guanine (G) in DNA, and uracil (U) in RNA – in a nucleic acid sequence. In sequencing assays, these repeats can cause errors or difficulties in sequence alignment and analysis.
Horizontal gene transfer
Horizontal gene transfer, also known as lateral gene transfer, is a process in which an organism transfers genetic material to another organism that is not its offspring. Horizontal gene transfers play a major role in bacterial evolution and can be divided into three different types: Transformation, conjugation and transduction.
Hot-start polymerase
During PCR set-up at room temperature, unintended products and primer dimers may be produced due to non-specific DNA amplification. Hot-start polymerases that remain inactive until the PCR reaction is heated have been developed to solve this problem and eliminate the need for preparing PCR reactions on ice.
I
IC50
Half-maximal inhibitory concentration (IC50) is the most widely used and informative measure to indicate how much of a specific substance is needed to inhibit a biological or biochemical process by half. For example, IC50 is used in pharmacological research to provide a measure of the potency of an antagonist drug.
Illumina sequencing
Illumina sequencing is a synthesis method used to sequence millions of DNA fragments in parallel. For a standard Illumina protocol, a library is prepared by fragmenting, purifying and amplifying the DNA sample. The sequence of each of these fragments is then determined simultaneously, and computationally aligned against a 'normal reference' genome.
Immunoglobulins
See antibody.
Immunoprecipitation (IP)
The technique of using an antibody that binds to a particular protein to precipitate it out of a solution is called immunoprecipitation (IP). IP requires that the antibody is coupled to a solid substrate at some point in the procedure, and can be used to isolate and concentrate a specific protein from a sample containing thousands of different proteins.
L
Laboratory centrifuge
A laboratory centrifuge is a device with a central motor that spins a rotor containing sample tubes. The resulting centrifugal force speeds up the sedimentation process, and centrifuges are commonly used to separate particles with different sizes and densities suspended in a liquid.
Centrifuges designed to accommodate tubes with a capacity of 2 ml and less are called microcentrifuges, and instruments optimized for very high speed – generating forces of up to 1,000,000 g – are called ultracentrifuges.
Laminar flow hood
Laminar flow hoods (or laminar flow cabinets) are enclosed, ventilated workspaces equipped with high efficiency particulate air (HEPA) filters. They are designed to protect samples from aerosol contamination by directing filtered air over the work zone into the lab.
Library preparation
Liquid handling
Liquid handling means transferring liquid from one location to another. As there are often countless samples of only a few microliters in volume that need to be transferred, accuracy, precision and throughput are key.
Liquid handling platform
Loop-mediated isothermal amplification (LAMP)
Loop-mediated isothermal amplification (LAMP) is a technique used to amplify DNA. Unlike PCR reactions, it can be performed at a constant temperature, because a DNA polymerase with strand displacement activity eliminates the need for heat denaturation.
M
Magnetic bead purification
Magnetic bead-based purification methods use magnetic beads with functionalized silica surfaces to selectively bind nucleic acids (including genomic DNA, plasmids, RNA and mitochondrial DNA), proteins and other biomolecules, which can then be easily separated from the aqueous phase with a magnet.
MALDI-TOF
MALDI-TOF is a two-phase mass spectrometry procedure. In the initial step, samples fixed in a crystalline matrix are vaporized and ionized by a laser. High voltage is then applied to accelerate the charged particles, and a detector measures the time of flight of each particle, enabling it to determine their molecular masses.
Manual pipette
Manual pipettes (or mechanical pipettes) require manual operation of the piston by pushing the plunger down with the thumb. Pipette tips also need to be ejected manually via the tip ejector and the volume is usually adjusted by twisting the plunger.
Mechanical pipette
See manual pipette.
Media preparation
Media preparation refers to the process of mixing nutrients and other substances to produce a culture medium for the growth of microorganisms, such as bacteria or fungi.
Media sterilization
Media sterilization refers to the process of sterilizing a culture medium to provide a sterile growth environment for microorganisms, such as bacteria or fungi.
Metabolomics
Metabolomics is the study of metabolites, substrates and products of chemical reactions within cells. By determining the metabolites and their concentrations in cells and tissues, metabolomics can directly reflect underlying biochemical activity and cell state (chemical fingerprint).
Metagenomics
Metagenomics is a branch of genomics that involves the study of genetic material directly extracted from environmental samples – such as soil or seawater – instead of from a single organism. It provides a comprehensive picture of the microbial community in a specific environment, revealing information about species diversity and interactions.
Methylation
Methylation is a process where a methyl group (a carbon atom linked to 3 hydrogen atoms, CH3) is added to a molecule. In genetics, DNA methylation can affect gene expression without altering the DNA sequence. This process plays a crucial role in aging, age-related diseases and longevity.
Microbiome
Microbiome is a term that describes the genome of all the microorganisms, symbiotic and pathogenic, living in a particular habitat, e.g. the human gut. The human microbiome has extensive functions in development of immunity, defense against pathogens, synthesis of vitamins and fat storage, as well as an influence on human behavior.
Micropipette
Micropipettes are essential laboratory instruments used to accurately and precisely transfer volumes of liquid in the microliter range. They are available in mechanical and electronic, as well as in single- and multichannel variants.
Microplate
Microplates, also called microtiter plates, are multiwell plates employed for various life sciences applications, such as diagnostics, molecular biology, pharmaceutical development and biomedical studies. They feature multiple wells placed side by side in a rectangular matrix, and are available in a variety of sizes, including 24, 48, 96, 384 and 1536 well plates.
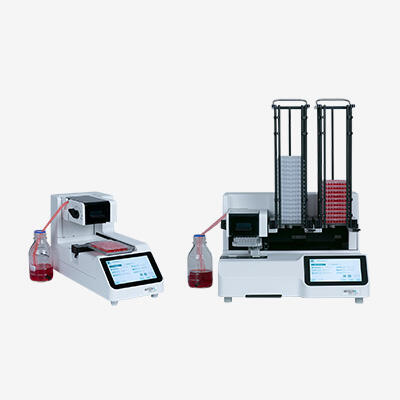
Microplate dispenser
Microplate dispensers are used to accurately and rapidly fill 6, 12, 24, 96, 384 and 1536 well plates by dispensing a predefined amount of liquid into each well. Microplate dispensers are non-contact dispensing systems that eject the liquid as a free-flying droplet or jet from the tip, and do not need to contact the microplate to break the surface tension of the liquid aliquot.
Microtiter plate (MTP)
See microplate.
Miniprep
Miniprep is a technique used to isolate plasmid DNA from bacteria for cloning or analysis, e.g. sequencing or restriction analysis.
MIQE guidelines
The MIQE (Minimum Information for Publication of Quantitative Real-Time PCR Experiments) guidelines describe the minimum information necessary for evaluating qPCR experiments. When publishing a manuscript, the scientist needs to provide all relevant experimental conditions and assay characteristics described by the MIQE guidelines, allowing reviewers to assess the validity of the protocols used, and enabling other scientists to reproduce the experiments.
Most probable number (MPN)
The most probable number (MPN) method is used to determine the concentration of a countable entity, e.g. viable microorganisms in a sample. To determine the concentration of the entity of interest, multiple serial dilutions of the sample are performed, the presence or absence of the entity in each replicate is assessed to, ultimately, calculate its concentration in the initial sample.
Multi-dispensing
See repeat dispensing.
Multi-resistant bacteria
Multi-resistant (or multidrug-resistant) bacteria are resistant to more than one antibiotic. Finding a treatment for an infection caused by multi-resistant bacteria can be difficult, and force physicians to prescribe antibiotic compounds with severe adverse effects. For example, when E. coli is resistant to ciprofloxacin and doesn't respond to a treatment with fluoroquinolones, colistin – an antibiotic that had gone out of use for adverse effects, such as kidney damage – often is the last resort treatment.
- https://www.reactgroup.org/toolbox/understand/antibiotic-resistance/multidrug-resistant-bacteria
- https://www.who.int/news-room/fact-sheets/detail/antimicrobial-resistance
- https://www.bag.admin.ch/bag/en/home/krankheiten/infektionskrankheiten-bekaempfen/antibiotikaresistenzen/wie-entwickelt-sich-die-antibiotikaresistenzlage---.html
Mutagenesis
The formation of mutations in DNA molecules is called mutagenesis. Such mutations can occur spontaneously in cells, be caused by genotoxic agents (e.g. radiation) or be produced using laboratory procedures.
N
Next generation sequencing
Next generation sequencing (NGS) is a technique used to sequence millions of DNA fragments in parallel. First, a library is prepared by fragmentation, purification and amplification of the DNA sample. Individual fragments are then physically isolated by attachment to solid surfaces or small beads. The sequence of each of these fragments is determined simultaneously, and computationally aligned against a ‘normal reference’ genome. This enables the detection of many sequence alterations, such as SNPs, in a single reaction.
Non-contact liquid dispensing
When dispensing liquids in the nanoliter or microliter range, contact dispensers need to contact the target (e.g. the test tube or microplate) to break the liquid aliquot's surface tension. In contrast, non-contact liquid dispensing systems, such as the VIAFILL, provide sufficient kinetic energy to eject the liquid from the tip as a free-flying droplet or jet.
Nuclease
Nucleases are enzymes cleaving the phosphodiester bonds of nucleic acid backbones. Nucleases degrading DNA are called deoxyribonucleases (DNases), and nucleases degrading RNA are called ribonucleases (RNases).
Nucleic acid extraction
To extract DNA, RNA or total nucleic acids from samples – such as blood, cultured cells, microbes, soil, or plant and animal tissues – the samples undergo a varying set of manipulations, depending on the sample and downstream application. These include cell lysis, inactivation of nucleases, and purification to separate the desired nucleic acid from cellular debris.
O
Organoid
An organoid is a 3D multicellular in vitro tissue construct that mimics its corresponding in vivo organ. Most commonly derived from stem cells – either pluripotent (embryonic or induced) or adult stem cells from various organs – organoids are used to study certain aspects of an organ.
P
Pan-resistant bacteria
Pan-resistant (or pandrug-resistant) bacteria are resistant to all currently known antibiotic compounds. This means that there is no effective antibiotic treatment for infections with pan-resistant bacteria at present.
Passaging cells
Passaging, or subculturing, refers to the process of splitting a cell culture into different vessels, and providing fresh medium to keep the cells at an optimal density for continued growth and proliferation. If adherent cells are not subcultured, they will eventually run out of room for expansion, while cells in suspension cultures will exceed the capacity of the medium to support further growth.
Pasteur pipette
Pasteur pipettes, also known as droppers, are plastic or glass tubes that are tapered to a narrow opening at the lower end, and fitted with a plastic or rubber bulb at the upper end. The tapered lower end makes them especially useful for transferring small volumes of liquid. Pasteur pipettes were invented by the famous 19th century scientist Louis Pasteur, and are still used for certain applications (e.g. titrations) in today's laboratories.
PCR
The polymerase chain reaction (PCR) is a fast and inexpensive technique for amplifying a DNA sequence of interest. It consists of three steps:
- Denaturation: The sample is heated to separate the DNA into two single strands.
- Annealing: The temperature is lowered to allow primers to anneal to specific single-stranded DNA segments, flanking the sequence to be amplified.
- Extension: The temperature is raised to the optimum working temperature of the polymerase enzyme, which then makes a complementary copy of the DNA sequence of interest.
One such repetition or 'thermal cycle' theoretically doubles the amount of the DNA sequence of interest present in the reaction. Typically, 25 to 40 cycles are performed – resulting in millions or even billions of DNA copies – depending largely on the amount of DNA in the starting sample and the number of amplicon copies needed for post-PCR applications.
PCR inhibitor
PCR inhibitors are substances preventing nucleic acid amplification by:
- binding to nucleic acids
- binding to polymerases
- binding essential co-factors like Mg2+
- interfering with nucleic acid replication
- preventing primers from annealing to nucleic acids
PCR-grade water
PCR-grade water is certified to be free of contaminants that may lead to false positive results, nucleases degrading nucleic acid samples and inhibitors that may interfere with polymerases or interact with the nucleic acid samples.
Peristaltic pump
A peristaltic pump operates by mechanically deforming flexible tubing (often silicone) to move liquid through it. This tube squeezing mechanism is called peristalsis, and is a common principle used by many biological systems, such as the gastrointestinal tract, for moving various fluids.
Petri dish filler
Petri dish fillers are systems used to fill Petri dishes with agar medium. Petri dish fillers eliminate the risk of contaminating the agar medium by removing the need for manual manipulations.
Pipette accuracy
Accuracy is the ability of a measuring instrument to give responses to a true value. A pipette is accurate to the degree that the volume delivered is equal to the specified volume.
Pipette aid
See pipette controller.
Pipette calibration
Calibration ensures that the difference between the selected and dispensed volume of a pipette is within the specifications. The most widely used calibration methods rely on gravimetric measurements. However, for instruments with 24, 96 or 384 channels, a hybrid gravimetric/photometric method is recommended in order to guarantee that outliers, such as defective channels, are detected.
Pipette controller
Pipette controllers (or pipette aids) attach to glass and plastic serological pipettes to allow the aspiration, measurement and dispensing of liquids. The pipetting speed is controlled by varying the finger pressure on the aspiration and dispensing buttons.
Pipette precision
Precision expresses the degree of reproducibility, or agreement between repeated measurements. If you perform multiple dispenses of a certain volume with a pipette, and it dispenses actual volumes very close to each other, the pipette is precise and your results are reproducible.
Pipette reproducibility
See pipette precision.
Pipetting programs
Many pipettes and pipetting robots offer pre-programmed, customizable pipetting programs, as well as the possibility to create step-based custom programs from scratch. Pipetting programs minimize human errors and reduce operator-to-operator variability, by ensuring that the protocol is executed the same way every time.
Pipetting robot
Plasmid
A plasmid is a small, circular, double-stranded DNA molecule found in bacterial cells and some eukaryotes. Plasmids are separate from the chromosomal DNA, and often carry genes associated with genetic advantages, such as antibiotic resistance. When a bacterium divides, all of the plasmids are copied, such that each daughter cell receives a copy of them, and they can also be passed between different bacterial cells through a process called conjugation.
Plasmid purification
Plate reformatting
The process of transferring samples from one microplate format to another, e.g. from a 96 well plate to a 384 well plate, is called reformatting.
Positive displacement pipette
In positive displacement pipettes, the piston is in direct contact with the sample, so is part of the disposable tip, not the pipette itself, to avoid cross-contamination. The aspiration process is similar to air displacement pipettes: the piston moves up, creating a partial vacuum that is filled by liquid.
Pre-wetting
The process of aspirating and dispensing the nominal volume three times after loading tips onto a pipette is called pre-wetting. This will equilibrate temperature differences and humidify the ‘dead’ air space inside the pipette and tip. Neglecting the pre-wetting procedure can result in the first few dispenses delivering lower volumes due to evaporation.
Precision
See pipette precision.
Primers
Primers – also called oligonucleotides or oligos – are short, single-stranded nucleic acids used in the initiation of DNA synthesis. During PCR reactions, they anneal to the plus and minus strands of the template DNA, flanking the sequence that needs to be amplified.
Protein crystallography
Protein crystallography is a technique used to determine the macromolecular structure of proteins. The protein of interest is crystalized by supersaturation before shining x-rays at the resulting crystals. The diffraction pattern allows the protein structure to be determined.
Protein extraction
See protein purification.
Protein purification
Protein purification (or protein extraction) is used to isolate one or a few proteins from a complex mixture, usually containing cells, tissues or whole organisms. Purification is vital for the characterization of the function, structure and interactions of the protein of interest. There are several protein purification methods and, depending on the approach, they take advantage of the different sizes, physico-chemical properties or binding affinities of proteins to separate them from each other.
Q
qPCR
qPCR – also called real-time PCR, quantitative PCR or quantitative real-time PCR – is a technique used to detect and measure the amplification of target DNA as it is produced. In contrast to conventional PCR reactions, qPCR requires a fluorescent intercalating dye or fluorescently-labeled probes, and a thermal cycler that can measure fluorescence and calculate the cycle threshold (Ct) value. Typically, the fluorescence intensity increases proportionately to the concentration of the PCR product being formed, measuring quantities of the target in real time.
R
Reagent
A reagent is a compound or mixture added to a system to cause a chemical reaction or test if a reaction occurs. A reagent may be used to find out whether or not a specific chemical substance is present, by causing a reaction to occur with it.
Reference gene
A reference gene is a gene whose expression is invariant under the experimental conditions. As it is continually transcribed, it can be used for qPCR normalization. Reference genes are also sometimes referred to as 'housekeeping genes' or 'control genes'.
Reformatting
See plate reformatting.
Relative centrifugal force (RCF)
The relative centrifugal force is the force acting on samples during centrifugation. It is indicated as a multiple of the gravitational field of Earth (g) and can be calculated as follows:
RCF = 11.2 x r x (RPM/1000)2
r: rotation radius (in cm)
RPM: rotational speed (revolutions per minute)
Repeat dispensing
Repeat dispensing, also called multi-dispensing, is the process of aspirating a larger volume of liquid, then dispensing multiple smaller aliquots of the same volume. It reduces the number of pipetting steps by performing several dispenses with a single aspiration, without the need to refill the tip after each action.
Reproducibility
See pipette precision.
Retrovirus
Retroviruses, like HIV, have a single-stranded RNA genome. Once a retrovirus has entered a host cell, the reverse transcriptase enzyme synthesizes complimentary DNA (cDNA) from the retroviral RNA genome. In the next step of the virus’s life cycle, the viral DNA enters the nucleus and integrates into the host cell's genome. The host cell then starts to produce new copies of viral RNA by transcribing and translating the viral genes, allowing retroviruses to replicate and infect other cells.
Reverse pipetting
Reverse pipetting is the process of aspirating slightly more than the target volume, then dispensing the desired volume, with the small excess staying in the tip after the dispense. This technique can improve reproducibility for non-aqueous solutions, by minimizing the influence of evaporation in the tip for volatile solutions, or the influence of liquid adhering to the inner wall of the tip for viscous solutions.
Reverse transcriptase
Reverse transcriptase is an enzyme that synthesizes cDNA from RNA. It is used, for example, by retroviruses like HIV to transform their viral RNA into cDNA, making it capable to integrate into the host genome. Scientists use reverse transcriptase in the lab for applications such as RT-PCR to transcribe RNA into DNA before amplification.
RNA isolation
RNase
See nuclease.
RT-PCR
Reverse transcription PCR (RT-PCR) is used to amplify RNA target sequences, such as messenger RNA or RNA virus genomes. This type of PCR involves an initial incubation of the RNA samples with a reverse transcriptase enzyme and a DNA primer – comprising sequence-specific oligo (dT)s or random hexamers – prior to the PCR amplification.
S
Sanger sequencing
Sanger sequencing, also called the chain termination method, is a method used to determine the nucleotide sequence of DNA. The basic principle of Sanger sequencing is to synthesize a new strand of DNA complementary to the template strand, while incorporating modified, labeled nucleotides that can terminate the synthesis process. Sanger sequencing can be broken down into three steps: chain termination thermal cycling, size separation and determination of the DNA sequence.
SARS-CoV-2
See COVID-19.
SDS-PAGE
Sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) is an analytical method to separate proteins based on their molecular weight. The combined use of sodium dodecyl sulfate (also known as sodium lauryl sulfate) and polyacrylamide gel allow the influence of structure and charge to be eliminated, and proteins to be separated solely based on polypeptide chain length.
Serial dilution
A serial dilution is a step-wise series of dilutions, where the dilution factor stays the same for each step. The purpose of a serial dilution is to estimate the concentration of a sample, or to obtain the desired concentration of a reagent, chemical or compound.
Serological pipette
Serological pipettes are made from glass or plastic and feature a volume graduation. To aspirate and dispense liquid with a serological pipette, it needs to be attached to a rubber bulb or pipette controller.
Serotype
Serotypes are groups within a single species of microorganisms, such as bacteria or viruses, which share distinctive surface structures. Salmonella bacteria can for example be divided into more than 2500 serotypes, based on their distinct combination of O antigens located on the outer membrane surface and H antigens that are part of the flagella.
Solid-phase extraction
Solid-phase extraction is a sample preparation method used to separate dissolved or suspended compounds in a liquid mixture depending on their physical and chemical properties. The sample is usually passed through a cartridge containing a solid adsorbent that either retains the analyte or the impurities.
Spacer pipette
Spacer pipettes, also called adjustable tip spacing pipettes, enable the parallel transfer of multiple samples between labware of different sizes and formats, e.g. from 96 to 384 well plates or from tubes to plates.
Spectrophotometry
Spectrophotometry measures the proportion of light of a certain wavelength that is absorbed by a sample. The absorbance values allow both qualitative and quantitative analysis of the sample, e.g. by determining the purity of a solution or the concentration of a certain analyte.
Spheroids
Multicellular spheroids are three-dimensional, spherical multicellular aggregates and one of the most common and versatile ways to culture cells in 3D. Examples include multicellular tumor spheroids (MCTS), neurospheres, mammospheres, hepatospheres and embryoid bodies.
Sterility
Sterility is the state of being free from living organisms. Sterile products, such as pipette tips, are needed to prevent contamination in cell culture assays. For very sensitive applications, products should not only be sterile, but also free from DNase, RNase, PCR inhibitors and endotoxins.
Subculturing cells
See passaging cells.
T
Tip immersion depth
The tip immersion depth indicates how far the pipette tip is immersed into the liquid. It is best to immerse it just below the liquid's surface (2-3 mm), to reduce the risk of liquid droplets clinging to the outside of the tip.
Titration
Titration is a chemical analysis method used to determine the concentration of an unknown solution. To calculate its concentration, a solution of a known concentration that reacts in a definite, known proportion with the unknown solution is progressively added until the reaction neutralizes.
Touch off
After dispensing liquid with a pipette, a droplet of residual liquid often clings to the tip end. You should remove it by performing a dry touch off (on the sidewall of the vessel or on the liquid surface in the container) or a wet touch off (directly dispense into the liquid).
Toxins
Toxins are substances produced by a living organism that are harmful to some other organisms. Toxins such as animal or plant venoms usually consist of a complex mixture of substances that together exert a greater effect than a single substance. Toxins can also be produced by microorganisms such as bacteria, viruses or fungi.
Transcriptome
The transcriptome represents the complete set of RNA molecules – including mRNA, rRNA, tRNA, and other non-coding RNAs – present in a cell. Studying the transcriptome can help understand gene structure, expression and regulation.
V
Volume range
The volume range indicates the maximum and minimum quantity of liquid that a pipette can transfer. As a rule of thumb, always choose the smallest pipette capable of handling the required volume, as its accuracy decreases close to its minimum capacity.
vPCR
For viability PCR (vPCR), each sample needs to be split into two aliquots. One aliquot is incubated with a photoreactive intercalating dye that is unable to diffuse through intact cell membranes of live cells. This means that it only intercalates into the DNA of dead cells. When this aliquot is subsequently treated with a blue light, the dye binds irreversibly to the DNA. Both aliquots are then subject to DNA purification and qPCR amplification. If they exhibit similar qPCR signals, the target microorganisms in the sample are mostly viable. If the dye-treated aliquot exhibits a weaker signal, the target microorganisms are mostly dead. vPCR is an important technique in diagnostics, agriculture and food safety.
W
Western blotting
Western blotting is a technique used to detect the presence of a specific protein in a complex mixture extracted from cells or tissues. This technique has four stages: the separation of proteins by size using gel electrophoresis, the transfer of separated proteins to a solid support, the detection of a target protein by antibodies, and the visualization of the target protein as a band on a blotting membrane using an imaging system.
Wet touch off
See touch off.
Y
Yeast
Yeasts are eukaryotic, unicellular fungi reproducing asexually by budding or fission, and sexually by spore formation. Around 1,000 yeast species are known, but this represents only a fraction of yeast biodiversity on Earth.