Setting up a PCR lab from scratch
Written by Stephanie Stursberg
10. June 2021
Setting up a new lab is exciting, but can also be a daunting process. We've recently faced this challenge ourselves, so we’ve put together our learnings in this five-part series. After discussing what is needed for nucleic acid extraction in part two, we'll now take a look at what needs to be considered when performing polymerase chain reactions (PCRs).
Table of contents
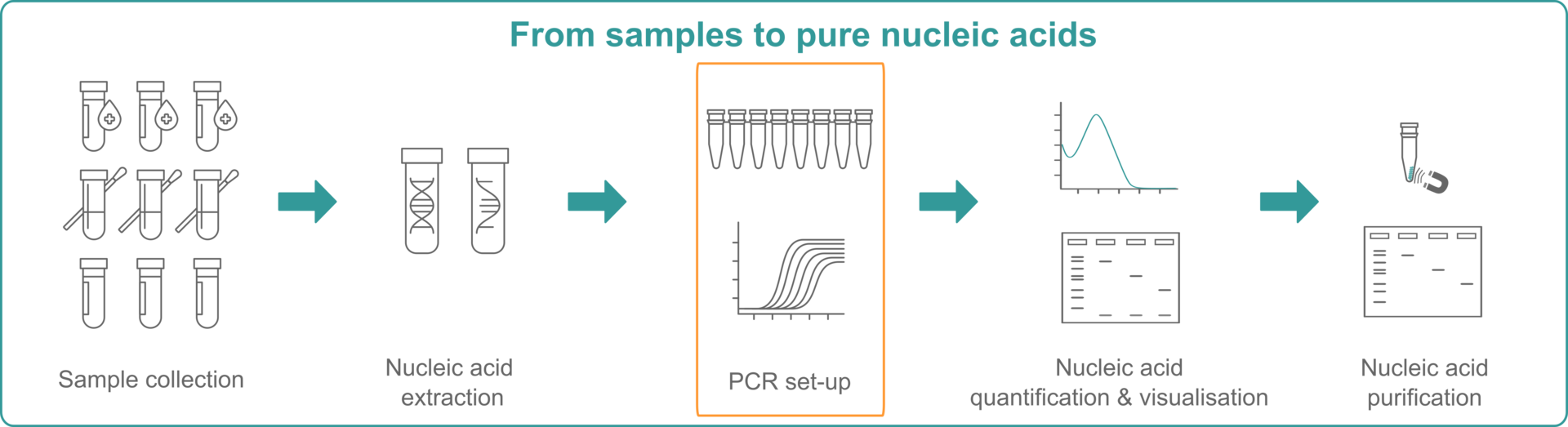
1 How do PCR, qPCR and RT-PCR work?
Real-time and reverse transcription PCR are both often abbreviated to ‘RT-PCR’ but they are not the same thing.1 To make sure that we're all on the same page, we will briefly outline the differences between these abbreviations – PCR, qPCR and RT-PCR – before exploring each technique further:
PCR
The polymerase chain reaction (PCR) is a fast and inexpensive technique for amplifying a DNA sequence of interest. It consists of three steps:
- Denaturation: The sample is heated to separate the DNA into two single strands.
- Annealing: The temperature is lowered to allow primers to anneal to specific single-stranded DNA segments, flanking the sequence to be amplified.
- Extension: The temperature is raised to the optimum working temperature of the polymerase enzyme, which then makes a complementary copy of the DNA sequence of interest.
One such repetition or 'thermal cycle' theoretically doubles the amount of the DNA sequence of interest present in the reaction. Typically, 25 to 40 cycles are performed – resulting in millions or even billions of DNA copies – depending largely on the amount of DNA in the starting sample and the number of amplicon copies needed for post-PCR applications.
Interesting fact: The polymerase enzyme typically used in PCRs is called Taq-polymerase, because it was originally isolated from the bacterium Thermus aquaticus. First discovered in a hot spring in Yellowstone National Park in the 1960s, this enzyme can resist the high temperatures necessary for DNA denaturation.
qPCR
qPCR – also called real-time PCR, quantitative PCR or quantitative real-time PCR – is a technique used to detect and measure the amplification of target DNA as it is produced. In contrast to conventional PCR reactions, qPCR requires a fluorescent intercalating dye or fluorescently-labeled probes, and a thermal cycler that can measure fluorescence and calculate the cycle threshold (Ct) value. Typically, the fluorescence intensity increases proportionately to the concentration of the PCR product being formed, measuring quantities of the target in real time.
RT-PCR
Reverse transcription PCR (RT-PCR) is used to amplify RNA target sequences, such as messenger RNA or RNA virus genomes. This type of PCR involves an initial incubation of the RNA samples with a reverse transcriptase enzyme and a DNA primer – comprising sequence-specific oligo (dT)s or random hexamers – prior to the PCR amplification.
PCR reactions are very sensitive and create a large number of copies of nucleic acids from minute amounts of starting material. This makes them a fundamental and highly effective molecular biology technique. However, because it is prone to amplicon and sample contamination, planning and designing of your PCR lab space will need careful consideration.
2 Designing your PCR lab
Ideally, a PCR lab should have two rooms with two areas, each designed for specific tasks. The first room should be exclusively used for pre-PCR activities and divided into a master mix preparation area and a sample preparation area. Air pressure should be slightly positive to prevent aerosols from flowing in.
The second room should have a dedicated area for nucleic acid amplification, and another one for product analysis. Air pressure should be slightly negative to ensure that amplicon aerosols don't leave the room.
If you're lacking in space or budget for a two-room PCR lab, you can set up the pre-PCR and amplification and analysis areas in the same room, but ensure they are as far from one another as possible.
Having pre-PCR activities spatially separated from the amplification and analysis area – either in different rooms or in separate benches – is very important, because you usually have a low amount of the nucleic acid sample during preparation and a very high concentration after amplification. This means that if you analyze your PCR in the same space as you prepare your master mix and samples, you may get false-positive results due to amplicon contamination.
You should also ensure that your lab set-up follows a unidirectional workflow. No materials or reagents used in the amplification and analysis areas should ever be taken into the pre-PCR space without a thorough decontamination. This means that you'll need dedicated equipment for each area, e.g., two different sets of pipettes. This unidirectional workflow should also apply to lab staff. If you've been working in the amplification and analysis areas, and you need to go back to the pre-PCR area, change your personal protective equipment, as it may have been contaminated by amplicon aerosols.
Another precautionary measure to take into account when setting up your PCR lab, in addition to the spatial separation, is temporal separation. You could, for example, consider setting up your PCR reactions in the morning, and perform the amplification and analysis steps in the afternoon. This may limit your flexibility, but will prevent contamination issues and having to repeat your experiment.
3 PCR equipment tips
PCR labs typically require a variety of equipment, such as centrifuges, vortex mixers, pipettes, fridges and freezers, thermal cyclers and analysis instruments (e.g., electrophoresis systems). Depending on the size of your lab and your applications, the amount of equipment you’ll need may vary. Instead of providing you a 'shopping list', we will outline what you should look for when purchasing equipment and consumables in order to keep contamination of your PCR reactions to a minimum.
3.1 Laminar flow or biosafety cabinet
Since you can never be 100 % certain that there are no amplicon aerosols in your pre-PCR space, you should set up your PCR reactions in a laminar flow hood or biosafety cabinet, decontaminated with a bleach solution prior to starting and after you finish your work.
3.2 Pipette tips and other consumables
Despite being more expensive than normal pipette tips, using filter tips for your PCR set-up will avoid aerosols entering and contaminating your pipette, and avoid aerosols that might already be present in your pipette contaminating your master mix or samples. To minimize your filter tip consumption, first fill all your tubes with the master mix using only one tip or set of tips – if you're using multichannel pipettes – and follow with your samples, using one tip per sample. Adding the sample last is also recommended because it's easier to dispense it into a liquid than into an empty tube, and because it reduces the risk of aerosolizing your sample as you pipette. Learn the best pipetting practices to help prevent aerosol formation.
For consumables, you should make sure that you have enough small vials available in your lab when your PCR reagents arrive. Aliquoting them into smaller containers will increase their shelf life and prevent them from going through too many freeze/thaw rounds. If your reagents get contaminated, it will also save you from throwing away your entire supply, as you’ll have clean aliquots available for a second PCR.
Finally, you’ll need to make sure that all consumables and equipment are free of DNase, RNase and PCR inhibitors. Always chose sterile products from manufacturers that can certify that their tips and consumables are free of any of these potential contaminants.

4 Cleaning and contamination control
You won’t need to worry about cleaning or contamination control when setting up your lab, but you will when your lab is up and running. We will briefly address this topic below.
Whether you decide to set up your PCR reactions in a laminar flow hood, a biosafety cabinet or an open bench, you will need to decontaminate your work space before and after set-up by wiping it with a freshly made bleach solution and distilled water. The same process should be performed in the amplification and analysis areas. You should also make sure you clean your pipettes, equipment, doorknobs, and the handles of your fridges and freezers regularly.
Because PCR assays are so sensitive, all the preventative measures described here may still not guarantee that your experiments will never get contaminated. It is therefore necessary to include the appropriate controls to detect contamination early. Always include negative and positive controls, as this will help identifying master mix contaminations, and confirm the performance of the extraction protocol, reagents and amplification steps. Additionally, you should monitor the positivity rate in your lab, and ensure that unexpected increases in detection have identifiable causes, e.g., a seasonal outbreak.
5 Conclusions
We hope that you are still just as excited about setting up your PCR lab, and that this blog has made the task less daunting for you. If you're curious to what you'll need for subsequent nucleic acid quantification workflows, read part four of our molecular biology lab set-up series.
Further reading: