Improving the reproducibility of cell culture handling
Sources of variation during handling
Stress
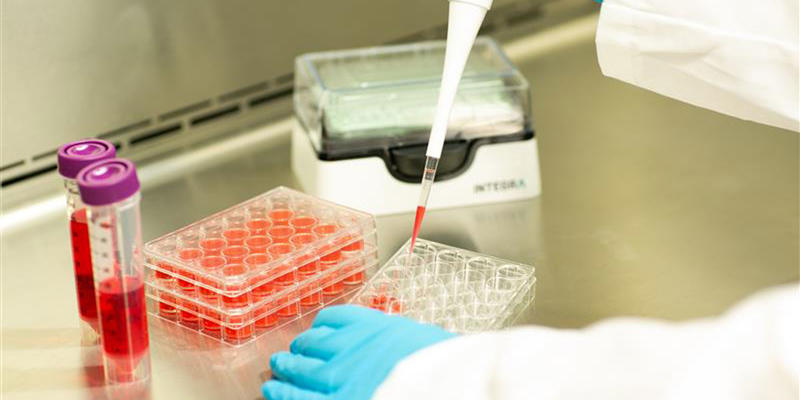
Cell culture parameters are routinely monitored to provide reproducible and viable cells, but variations in handling techniques are often overlooked. An analysis of possible variations in cancer screening applications found that results were far more consistent within a research group than between groups, with the latter observing a 200-fold variation in growth inhibition rates.2 This is likely down to differences in experimental methods, pipetting techniques and quality of equipment between the different research teams. Each of these factors subjects cells to different levels of stress, and can result in variations in cell viability, as well as decrease the reproducibility of experiments. Shearing forces are an unavoidable source of stress that cells encounter during pipetting, particularly if they are forced through a pipette tip that is too narrow or at a rate that is too fast. This action can lead to phenotypic and behavioral changes and, ultimately, result in variable cell viability. Therefore, a lab group using wide bore tips may obtain higher cell viability and more accurate conclusions than one using standard bore tips for cell culture.
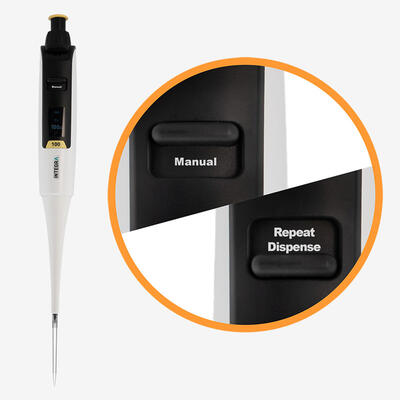
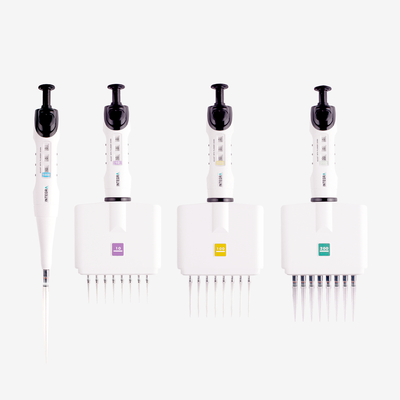
When working with manual pipettes, the pipetting angle will inevitably vary from user to user, as will the speed of aspiration and dispensing – depending on how quickly the plunger is moved – creating further variations in stress levels. Sudden starts and stops, or jerky movements, can introduce inconsistencies that go unnoticed if not well documented by researchers at the time. Electronic pipettes allow pipetting and mixing speeds to be clearly defined and documented, minimizing differences in technique between users or pipetting steps. Combining wide bore tips with electronic pipettes can give all users greater control over bioprocessing forces, helping to improve cell viability.
Deviating from protocols
For the reproducible growth of cells, researchers need to stick as close to the outlined protocol as possible, as any deviation can lead to inconsistencies. For example, variations in the number of cells seeded, or the distribution of cells in the medium, can cause variable cell growth, impacting results. For cytotoxicity assays, for example, the number of cells can significantly increase the minimum inhibitory concentration of an antibiotic – a phenomenon called the ‘inoculum effect’ – so cell numbers need to be consistent. Frequent mixing of the medium ensures that cells are suspended and equally distributed in the solution, preventing the formation of a cell concentration gradient between the bottom and the top of the vessel. This action needs to be consistent across experiments for the equal distribution and seeding of cells. However, mixing is exerting a shearing force over the cells that may result in stress, so overmixing should also be avoided to prevent variations in cell viability. Sticking as close to the protocol as possible – and accurately reporting all methodologies, mixing steps and pipetting speeds – can help to improve the reproducibility of cell handling tasks.
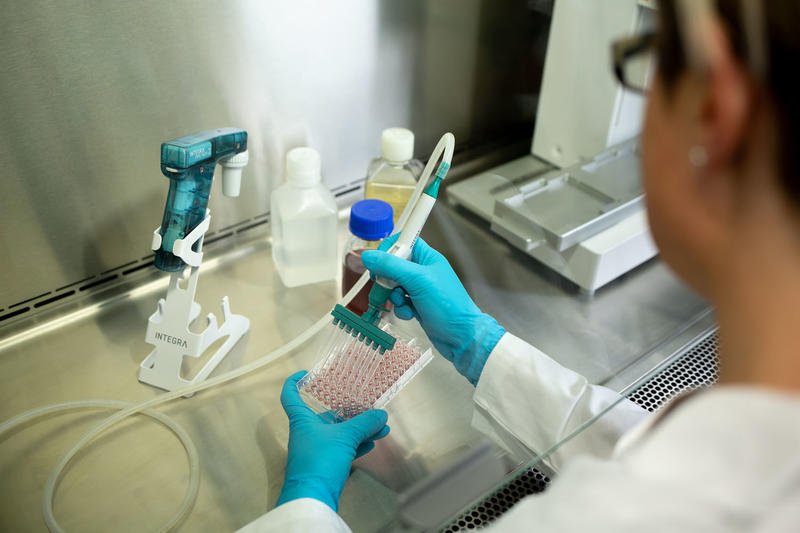

Contamination
Biological contamination of samples by microorganisms, and the cross-contamination between different cell lines during cell handling, can impact the health of cells and produce unreliable results. If undetected, contamination can lead to false conclusions about the survivability and behavior of cells, compromising the integrity and reproducibility of research. The correct handling of cell samples, medium and reagents should include aseptic procedures and equipment – such as sterile pipettes and tips, use of filter tips and workflow automation – to prevent the contamination of samples, along with effective quality control testing for unwanted microorganisms.
Ensuring reproducible handling
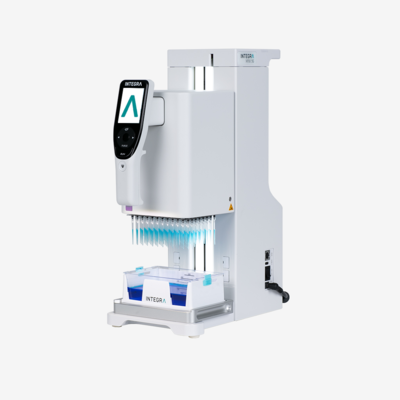
Automation of workflows can be beneficial for improving reproducibility and controlling the shearing forces acting on cells, eliminating the unavoidable variability associated with manual processing. This is particularly important when working with a large throughput of samples in 384 well plates, which is almost impossible to achieve manually without a higher rate of inconsistencies. Switching to automated processes can help to overcome this variability, providing greater control and uniformity of bioprocessing forces and cell manipulations.3
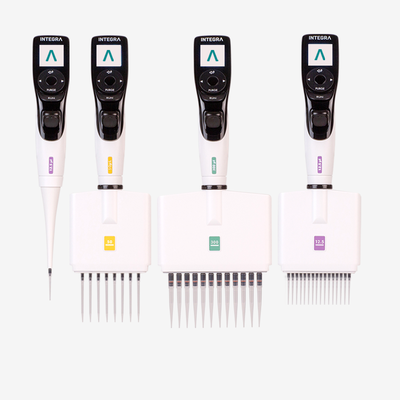
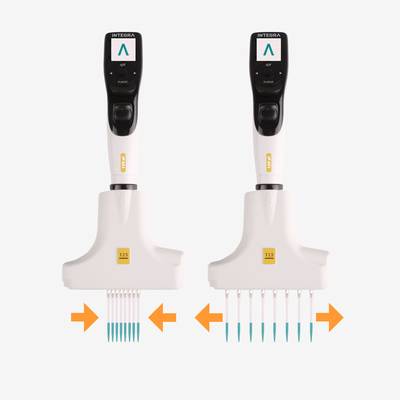
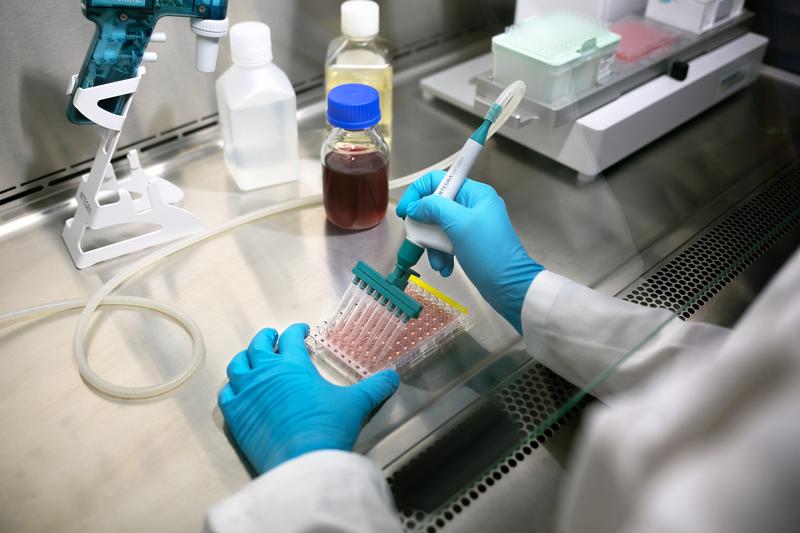
Ideally, every pipetting stage in cell culture should be automated – from plating to treatment – to reduce user-induced variation, and electronic pipettes provide a simple and affordable place to start. With an electronic pipette, users can select, document and save pipetting protocols with defined dispense speeds and mixing routines, so that each action is performed at the same rate every time, regardless of user experience or technique. In addition, repeat dispense modes enable a large volume of liquid to be aspirated and dispensed in small, defined volumes, reducing the need to move back and forth between the source and the plate, which can introduce inconsistencies and handling errors. Along with electronic pipettes, all plastic labware – including pipette tips – should be consistent, as deviations in the tip shape and dimensions, as well as the orifice diameter, may lead to different effects on the cells.
Going one step further and fully automating the entire pipetting workflow using a robot should eliminate the influence of user technique or experience altogether. Although there is likely to be a learning curve for the transfer phase from manual techniques to automated processes – to familiarize staff with new equipment and supplies – a hands-off workflow ultimately removes the user as a hidden source of variability and contamination, increasing the reproducibility of handling techniques.
- Baker, M (2016). Reproducibility crisis. Nature, 533(26), 353-66.
- Niepel, M., et al. (2019). A multi-center study on the reproducibility of drug-response assays in mammalian cell lines. Cell systems, 9(1), 35-48
- Brindley, D., et al. (2011). Bioprocess forces and their impact on cell behavior: implications for bone regeneration therapy. Journal of tissue engineering.